RIBOTAC technology may start a new era of RNA degradation
- EPA Announces First-Ever Regulation for “Forever Chemicals” in Drinking Water
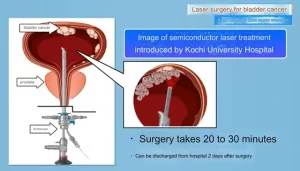
- Kochi University pioneers outpatient bladder cancer treatment using semiconductor lasers
- ASPEN 2024: Nutritional Therapy Strategies for Cancer and Critically Ill Patients
- Which lung cancer patients can benefit from neoadjuvant immunotherapy?
- Heme Iron Absorption: Why Meat Matters for Women’s Iron Needs
- “Miracle Weight-loss Drug” Semaglutide Is Not Always Effective
RIBOTAC technology may start a new era of RNA degradation
- Red Yeast Rice Scare Grips Japan: Over 114 Hospitalized and 5 Deaths
- Long COVID Brain Fog: Blood-Brain Barrier Damage and Persistent Inflammation
- FDA has mandated a top-level black box warning for all marketed CAR-T therapies
- Can people with high blood pressure eat peanuts?
- What is the difference between dopamine and dobutamine?
- How long can the patient live after heart stent surgery?
RIBOTAC technology may start a new era of RNA degradation.
PROTAC is called Proteolysis-Targeting Chimeras, or proteolysis-targeting chimeras.
It is an emerging drug type that is different from antibodies and traditional small molecule inhibitors.
After rapid development in recent years, the PROTAC field has made significant progress.
At present, many projects have entered the clinical development stage, which has brought unprecedented opportunities to solve the drug resistance of traditional small molecules and target non-drugable target proteins.
Similar to the PROTAC technology for targeted degradation of proteins, RIBOTAC technology for targeted degradation of RNA is also emerging.
The full name of RIBOTAC is Ribonuclease-Targeting Chimeras, that is, ribonuclease targeting chimeras, whose function is to “bring” target RNA to RNase for degradation.
On October 27th, in a study published in the journal Science Translational Medicine, RIBOTAC technology pioneer and Professor Matthew Disney of the Scripps Institute led a team to design a degradable amyotrophic lateral sclerosis (ALS) and frontal The RIBOTAC molecule of pathogenic RNA in temporal lobe dementia (FTD) [1] .
In both the patient-derived spinal cord neurons and ALS mouse models, RIBOTAC molecules successfully induced the degradation of pathogenic mRNA and reduced related pathology.
In the past 15 years, Professor Disney has been committed to studying the druggable structures of RNA molecules and has established a library of compounds that can combine these druggable structures.
The successful development of RIBOTAC is built on these foundations.
RIBOTAC first attracted attention in 2019, when Professor Disney published a key result in Cell Chemical Biology, describing their development of a RIBOTAC molecule that has anti-cancer potential and mediates selective RNA degradation [2] .
The development of RIBOTAC is based on small molecules that can selectively bind RNA (especially RNA that can form complex secondary and tertiary structures).
The key innovation lies in the transformation of RNA binding molecules into RNA binding molecules by linking ribonuclease (RNase) recruiting molecules. RNA degrading molecules.

Comparison of RIBOTAC and PROTAC | (A) RIBOTAC is a bivalent molecule that contains an RNA binding module (green), a ribonuclease (RNase) recruiting module (orange) and a linker (black line) that acts as a connection; Once bound to the target RNA, RIBOTAC will recruit RNase near the target RNA, thereby promoting its degradation. (B) PROTAC is also a bivalent molecule, in which the protein binding ligand (red) is connected to the E3 ligase recruitment module (black) through the linker (black line). PROTAC locates the E3 ligase near the target protein, leading to polyubiquitination of the target protein, and finally the target protein is degraded by the cellular proteasome mechanism. (Source: Cell Chemical Biology)
Specifically, this Cell Chemical Biology paper describes the RIBOTAC molecule TGP-210-RL that can mediate the selective degradation of pre-miR-210.
TGP-210-RL is derived from TGP-210, which is a small molecule that can bind pre-miR-210 previously developed by the Disney team.
TGP-210 is connected with tetra-adenylate to form TGP-210-RL. Tetra-adenylate is similar to the endogenous oligoadenylates (oligoadenylates) produced in cells during virus infection.
Oligoadenylate activates latent RNase L, dimerizes it and forms active ribonuclease.
Pre-miR-210 is an attractive therapeutic target because it is the precursor of microRNA miR-210, and microRNA miR-210 is essential for cancer survival in a hypoxic environment.
In this study, Professor Disney and his collaborators confirmed that TGP-210-RL can induce the formation of a ternary complex of TGP-210-R + microRNA precursor + RNase L.
In hypoxic cancer cells, TGP-210-RL induced a significant reduction in miR-210 levels and apoptosis, proving the potential of RIBOTAC to target and degrade RNA targets in cancer cells.
RIBOTAC’s second important paper was published in the Journal of the American Chemical Society (JACS) in August this year [3].
In this study, the scientists transformed an RTK inhibitor dovitinib into a small molecule RIBOTAC that targets RNA degradation.
Studies have confirmed that RIBOTAC, which targets the degradation of pre-miR-21 (the precursor of microRNA-21), alleviates disease progression in mouse models of triple-negative breast cancer and Alport syndrome (hereditary nephritis).
The RIBOTAC molecule modified based on dovitinib has increased selectivity for pre-miR-21 by 2500 times.


Source: JACS
In the latest study published in the journal Science Translational Medicine, Professor Disney’s team focused on neurodegenerative diseases ALS and FTD.
Both ALS and FTD are progressive neurodegenerative diseases that can lead to motor and cognitive impairment.
These diseases are usually sporadic, and the most common associated mutation is the hexanucleotide repeat amplification (HRE; GGGGCC repeat amplification) of an intron (usually intron 1) in the open reading frame 72 (C9orf72) of chromosome 9 , The number of repetitions that cause the disease ranges from 60 to thousands).
This subtype of the disease is called c9ALS/FTD.
RNA containing HRE is translated into a protein containing different dipeptide repeats (usually poly(GP) or poly(GA)). poly(GP) is polyglycine-proline, poly(GA) is polyglycine-alanine. Both HRE-containing RNA and its translational protein containing dipeptide repeats are thought to promote neuronal death.
Such RNA and proteins can form toxic aggregates, and RNA can block gene expression mechanisms.
Based on these early findings, scientists believe that if the RNA containing HRE can be eliminated, it may bring new opportunities for the treatment of diseases.
Compared with targeting proteins, targeting such pathogenic RNAs has the effect of “two birds with one stone”.
HRE-containing RNA has another feature worth targeting: this type of RNA will form a special 3D structure. Targeting this special 3D structure (rather than the primary sequence) may reduce targeting other shorter HREs. The off-target effect produced by the non-pathogenic mRNA.
Using the structure-activity relationship combined with biophysics and structural analysis, Professor Disney and his colleagues first designed a small molecule dimer compound that can bind to the above-mentioned special 3D RNA structure.
Each monomer is incorporated in the inner loop of the RNA hairpin. The binding Kd of the dimer is 4±0.7 nM, and it stays on the target RNA for a long time.

Design of RIBOTAC| (A) Compound 6 is a dimeric RNA binding module, and RNase L recruitment module is not connected; (B) Compound 7 is a RIBOTAC molecule coupled with RNase L recruitment module, and Compound 8 is coupled with a control, The less efficient RNase L recruitment module. (Source: Science Translational Medicine)
Then, on the basis of the dimer, Professor Disney’s team added a ribonuclease (RNase L) L recruitment module to turn the small molecule dimer into a RIBOTAC molecule. In HEK293T cells, their RIBOTAC molecule inhibited translation and dose-dependently reduced HRE-containing RNA levels in a way that required RNase L.
In addition, RIBOTAC molecules also alleviate the pathological characteristics of c9ALS/FTD patient-derived cell lines, and reduce the abundance of C9orf72 intron 1 in patient-derived lymphoblast cell lines and iPSCs, and the level of dipeptide repeat protein in iPSCs .
The same reduction can be seen in iPSC-derived spinal cord neurons (iPSNs, which can reproduce many genetic, transcriptional, and biochemical characteristics of brain tissue in c9ALS/FTD patients).
The study did not observe the effect of the designed RIBOTAC molecule on other transcripts.
Since a large number of previous studies have pointed out that in patients with c9ALS/FTD, nucleoporins will be reduced, therefore, this study also investigated the effect of RIBOTAC treatment on nucleoporins levels.
Through super-resolution microscopy, Professor Disney and others found that RIBOTAC restored the level of the key nucleoporin Nup98.

In preclinical mouse models, r(G4C2)exp targeting RIBOTAC 7 can reduce c9ALS/ftd-related toxicity (Source: Science Translational Medicine)
In the c9ALS/FTD mouse model, a single intracerebroventricular injection of RIBOTAC greatly reduced the levels of r(G4C2)-containing mRNA, r(G4C2)-containing foci, and poly(GP)-containing proteins.
These effects can be observed 1 week after the injection, and lasted until at least 6 weeks after treatment (that is, the entire study period). Through immunohistochemical analysis, RIBOTAC also reduced the known characteristics of c9ALS/FTD, including poly(GP) and poly(GA) aggregates, and the cytoplasmic content of TDP-43.
Scientists believe that although the advancement of RIBOTAC to the clinic requires further optimization of its pharmacochemical and physicochemical properties (this process may take several years), this new result provides another key evidence that the targeted degradation of RNA is It is feasible and may be the best way to treat diseases where RNA plays a key pathogenic role.
In terms of clinical translation, as early as 2016, Professor Disney founded the biotechnology company Expansion Therapeutics as the scientific founder, dedicated to the development of small molecule drugs for the treatment of RNA-mediated diseases.
In September of this year, Expansion just completed $80 million in Series B financing, and giants Novartis and Sanofi continue to support it.
Expansion currently focuses on neurodegenerative diseases, including the aforementioned c9ALS/FTD.
Summary
A variety of RNAs, including mRNA, miRNA, and long-chain non-coding RNA, are all related to diseases, so they are potential drug targets.
As a new type of molecule, RIBOTAC has the potential to target different types of RNA. It may have certain advantages over antisense oligonucleotides (ASO) and siRNA that have been developed to degrade RNA, including:
1) better drugs As a small molecule, RIBOTAC may enter a wider range of tissues, while ASO and siRNA have problems such as poor cellular uptake, poor tissue-specific delivery (except liver and kidney), and poor endosomal escape;
2) Lower Dosage: The catalytic properties of RIBOTAC are another major advantage. After RIBOTAC binds and induces degradation of one RNA, it can bind to another RNA.
In this way, a low concentration of RIBOTAC can be used to achieve the degradation of the target RNA.
However, RIBOTAC is also faced with challenges that are difficult to develop. The most difficult part is the development of small molecules that selectively bind to target RNA in cells.
Compared with antisense oligonucleotides, it is much more difficult to develop suitable RNA binding ligands, because The former can be simply programmed to bind to the target RNA using complementary sequences.
In addition, RIBOTAC also faces the challenges of early PROTAC, that is, poor cell absorption and low bioavailability due to its large molecular weight (the molecular weight of TGP-210-RL is> 2000.
Compared with TGP-210, cell uptake is reduced by 50%. %).
The further iteration of RIBOTAC may involve the search for smaller RNase binding ligands. It is worth noting that there have been reports of small molecule activators of RNase L in the literature.
Reference:
[1]JESSICA A. BUSH et al. Ribonuclease recruitment using a small molecule reduced c9ALS/FTD r(G4C2) repeat expansionin vitro and in vivo ALS models. SCIENCE TRANSLATIONAL MEDICINE(2021).
[2] Matthew G. Costales et al. Targeted Degradation of a Hypoxia-Associated Non-coding RNA Enhances the Selectivity of a Small Molecule Interacting with RNA. Cell Chemical Biology(2019).
[3]Peiyuan Zhang et al. Reprogramming of Protein-Targeted Small-Molecule Medicines to RNA by Ribonuclease Recruitment.JACS(2021).
[4] Degrading disease-causing RNA in ALS (Source: Nature Reviews Drug Discovery)
[5] Potential treatment for ALS, frontotemporal dementia targets RNA (Source: Scripps Research Institute)
[6] RIBOTACs: Small Molecules Target RNA for Degradation (Source: Cell Chemical Biology)
[7] Small-molecule selectively destroys cancer-associated RNA (Source: C&EN)
[8] Neuron | Sun Shuying’s team reveals the mechanism by which the RNA helicase DDX3X regulates the translation of C9ORF72 (Source: BioArt)
[9] Expansion Therapeutics Raises $80 Million Series B Financing to Treat Neurodegenerative Diseases (Source: Expansion)
RIBOTAC technology may start a new era of RNA degradation
(source:internet, reference only)
Disclaimer of medicaltrend.org
Important Note: The information provided is for informational purposes only and should not be considered as medical advice.