Major progress: New mitochondrial base editor successfully developed!
- Did Cloud Seeding Unleash a Deluge in Dubai?
- Scientists Identify Gut Bacteria and Metabolites that Lower Diabetes Risk
- OpenAI’s Model Matches Doctors in Assessing Eye Conditions
- UK: A Smoke-Free Generation by Banning Sales to Those Born After 2009
- Deadly Mutation: A New Monkeypox Variant Emerges in the DRC
- EPA Announces First-Ever Regulation for “Forever Chemicals” in Drinking Water
Major progress: New mitochondrial base editor successfully developed!
- Red Yeast Rice Scare Grips Japan: Over 114 Hospitalized and 5 Deaths
- Long COVID Brain Fog: Blood-Brain Barrier Damage and Persistent Inflammation
- FDA has mandated a top-level black box warning for all marketed CAR-T therapies
- Can people with high blood pressure eat peanuts?
- What is the difference between dopamine and dobutamine?
- How long can the patient live after heart stent surgery?
Major progress: New mitochondrial base editor successfully developed! A New Era of Mitochondrial Genome Editing Comes.
In a new study, researchers from the Center for Genome Engineering at the Korea Institute for Basic Science (IBS) have developed a new gene-editing platform called transcription activator-like effector-associated dehydrogenase. effector-linked deaminase, TALED). TALED is a base editor capable of A→G base transitions in mitochondria.
The discovery is the culmination of decades of treating human genetic diseases, and TALED can be considered the last missing piece of the puzzle in gene editing technology.

From the identification of the first restriction enzyme in 1968, the invention of the polymerase chain reaction (PCR) in 1985, to the demonstration of CRISPR-mediated genome editing in 2013, every new breakthrough discovery in biotechnology has been Further enhancing our ability to manipulate DNA, the blueprint for life.
In particular, the recently developed genetic scissors called the CRISPR-Cas system allow us to perform comprehensive genome editing in living cells.
This opens up new possibilities for treating previously incurable genetic diseases by editing mutations in our genomes.
While gene editing has been largely successful in the nuclear genome, scientists have, however, been unsuccessful in editing mitochondria, which also have their own genomes. Mitochondria, the so-called “energy factories of the cell,” are tiny organelles in cells. Since it is an important organelle of energy metabolism, if the gene is mutated, it will cause serious genetic diseases related to energy metabolism.
“There are some extremely nasty genetic diseases that arise from defects in mitochondrial DNA. For example, Leber hereditary optic neuropathy (Leber hereditary optic neuropathy), which causes sudden blindness in both eyes,” explains Kim Jin-Soo, corresponding author of the paper and director of the IBS Center for Genome Engineering. optic neuropathy (LHON) is caused by a simple single point mutation in mitochondrial DNA.” Another mitochondrial gene-related disorder is mitochondrial encephalomyopathy with lactic acidosis and stroke-like episodes, MELAS), it slowly destroys the patient’s brain. Some studies even suggest that abnormalities in mitochondrial DNA may also be the cause of degenerative diseases such as Alzheimer’s disease and muscular dystrophy.
The mitochondrial genome is inherited maternally. Mitochondrial DNA has 90 known pathogenic point mutations, which occur in at least 1 in 5,000 people.
Many existing genome editing tools cannot be used due to limitations in delivery methods to mitochondria.
For example, the CRISPR-Cas platform is not suitable for editing such mutations in mitochondria, because guide RNAs (gRNAs) cannot access the organelle itself.
Kim added, “Another problem is that animal models for these mitochondrial diseases are scarce. This is because it is currently impossible to engineer the mitochondrial mutations needed to build animal models. The lack of animal models makes it very difficult to develop and test treatments for these diseases. .”
Therefore, reliable technology for editing mitochondrial DNA (mtDNA) is one of the last frontiers of genome engineering, which must be explored in order to conquer all known genetic diseases, and which many scientists around the world have been working on for years to make it a reality.
In 2020, a research team led by David R. Liu of the Broad Institute constructed a new base editor called DddA-derived cytosine base editor (DdCBE), C→T transitions can be performed in mitochondrial DNA.
This was achieved by building a new gene-editing technique called base editing, which converts one nucleotide base to another without damaging the DNA. However, this technique also has its limitations. It is not only limited to C→T transitions, but mostly to TC motifs, making it effectively a TC→TT converter.
This means it can only correct 9 (10%) of the 90 confirmed pathogenic mitochondrial point mutations. For a long time, the A→G transition of mitochondrial DNA was thought to be impossible.
Cho Sung-Ik, the first author of the paper, said, “We started thinking about how to overcome these limitations.
The result is that we were able to build a new gene editing platform called TALED, which can achieve A→G transitions. Our new base editor Greatly expands the scope of mitochondrial genome editing.
This could make a huge contribution not only to making disease models, but also to developing treatments.” Remarkably, the mere ability to perform A→G transitions in human mtDNA allows correction 39 out of 90 known pathogenic variants (=43%).
The authors constructed TALED by fusing together three distinct components.
The first component is the transcription activator-like effector (TALE), which is capable of targeting DNA sequences. The second component is TadA8e, an adenine deaminase that facilitates the A→G transition.
The third component is DddAtox, a cytosine deaminase that makes DNA more accessible to TadA8e.
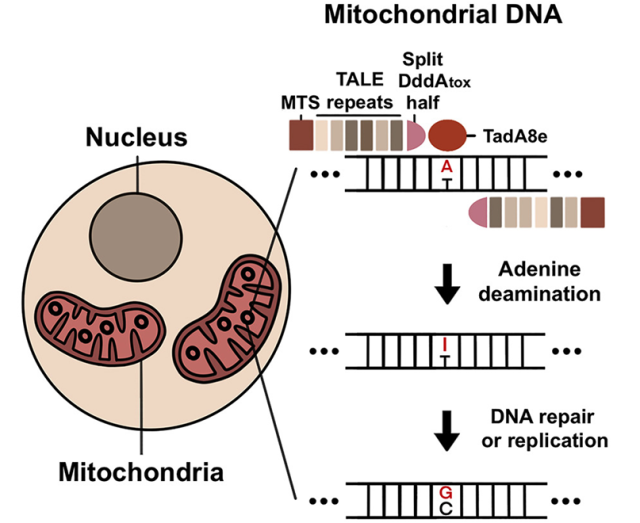
Image via Cell, 2022, doi:10.1016/j.cell.2022.03.039.
An interesting aspect of TALED is the ability of TadA8e to perform A→G editing in mitochondria with double-stranded DNA (dsDNA).
This is a mysterious phenomenon because TadA8e is a protein known to be specific only for single-stranded DNA (ssDNA).
“No one had thought of using TadA8e for base editing in mitochondria before, because it should only be specific for single-stranded DNA,” Kim said. “It was this out-of-the-box thinking that really helped us invent TALED.”
These authors speculate that DddAtox allows dsDNA to be accessed by transient unwinding of the duplex.
This fleeting but short-lived time window allows the super fast-acting TadA8e to quickly make the necessary edits.
In addition to tweaking the components of TALED, they developed a technique that enables both A→G and C→T base editing as well as A→G base editing only.
The authors demonstrated this new technique by creating single-cell-derived clones containing the desired mtDNA edits. Furthermore, they found that TALED was neither cytotoxic nor caused mtDNA destabilization.
Furthermore, there is no undesirable off-target editing in nuclear DNA, and few off-target effects in mtDNA.
Their goal is to further improve TALED by increasing editing efficiency and specificity, ultimately paving the way for correction of pathogenic mtDNA mutations in embryonic, fetal, neonatal or adult patients.
They are also focusing on developing TALEDs suitable for A→G base editing of chloroplast DNA. Chloroplast DNA encodes important genes in plant photosynthesis.
William I. Suh, IBS Science Communicator, praised, “I think the significance of this discovery is comparable to the invention of the blue LED, which won the Nobel Prize in 2014. Just like the blue light-emitting diode (LED) is what allows us to have high energy efficiency. Like the final piece of the puzzle for white LED light sources, TALEDs are expected to usher in a new era of genome engineering.”
References:
1. Sung-Ik Cho et al. Targeted A-to-G base editing in human mitochondrial DNA with programmable deaminases. Cell, 2022, doi:10.1016/j.cell.2022.03.039.
2. A new era of mitochondrial genome editing has begun
https://www.ibs.re.kr/cop/bbs/BBSMSTR_000000000738/selectBoardArticle.do?nttId=21220&pageIndex=1&searchCnd=&searchWrd=
Major progress: New mitochondrial base editor successfully developed!
(source:internet, reference only)
Disclaimer of medicaltrend.org
Important Note: The information provided is for informational purposes only and should not be considered as medical advice.