The role and expression of m6A in colorectal cancer
- Normal Liver Cells Found to Promote Cancer Metastasis to the Liver
- Nearly 80% Complete Remission: Breakthrough in ADC Anti-Tumor Treatment
- Vaccination Against Common Diseases May Prevent Dementia!
- New Alzheimer’s Disease (AD) Diagnosis and Staging Criteria
- Breakthrough in Alzheimer’s Disease: New Nasal Spray Halts Cognitive Decline by Targeting Toxic Protein
- Can the Tap Water at the Paris Olympics be Drunk Directly?
The role and expression of m6A in colorectal cancer
The role and expression of m6A in colorectal cancer. In order to evaluate the role of m6A in colorectal cancer, the author first evaluated the differential expression of m6A-related genes in colon cancer in the TCGA database.
Basic results of METTL3 in colon cancer
In order to evaluate the role of m6A in colorectal cancer, the author first evaluated the differential expression of m6A-related genes in colon cancer in the TCGA database. At the same time, they used samples from their own hospital to test the results of these differential expressions, and found that METTL3 There is a difference in expression among the verified results.
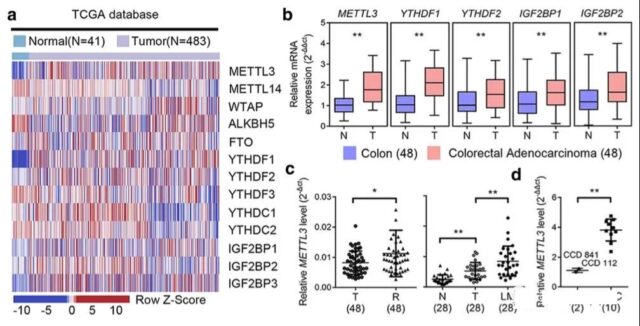
In addition to the analysis of cancer and normal tissue samples, the analysis of tissue samples will also further express whether there is any difference with various clinical information. So the author did an analysis on METTL3 and various clinical information. This generally will definitely do a lot of clinical data analysis, as for the results, it may be whichever is meaningful.
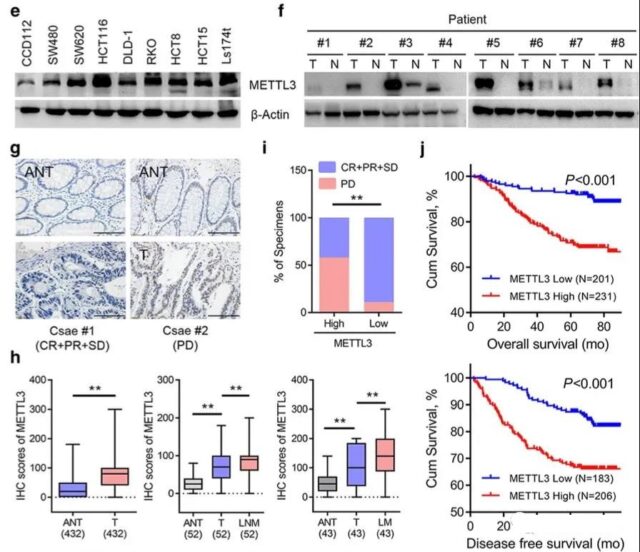
SOX2 is regulated by METTL3-mediated m6A methylation
When performing the expression verification of each cell line above, the author found that there are differences in the protein expression of METTL3 between SW480 and SW620. This pair of cell lines is a pair of cell lines isolated from the abdominal metastasis and primary tumor of a single patient. The two cell lines show different metastasis abilities, which may indicate the relationship between METTL3 and tumor metastasis.
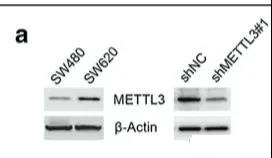
For m6A sequencing analysis, MeRIP-seq is used. So the author did two different MeRIP-seq analyses, one is SW480 vs SW620, and the other is SW620 vs SW620 cell line with METL3 knockout because of the high expression of METL3 in SW620 (this group of SW620 It’s probably the one in the first group before). Since we have also introduced that MeRIP-seq can only explain which position m6A is combined with, but it cannot explain the influence on expression. For the same grouping, the author also did an RNA-seq analysis.
Through the final crossover analysis, we can finally get that 158 genes are the result of two overlapping groups.
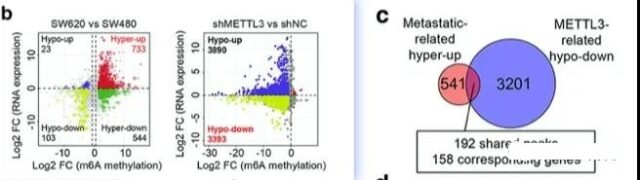
Further enrichment analysis of these 158 genes showed that these 158 genes are mainly related to the differentiation of stem cells, indicating that the transfer mediated by METL3 may be formally acting through the stem cell analysis pathway. The enrichment analysis website used by the author here is: metascape. There are many sites for enrichment analysis, and you can also use the WebSestalt enrichment analysis software we 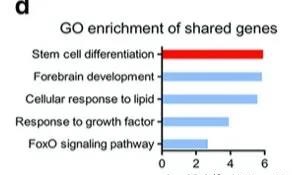
Now that you know that the stem cells are differentiated, it depends on the descriptive genes. So the author checked the analysis results and found that there are three genes, SEMA3A, BCHE, ZFP36L2 and SOX2 in the stem cell differentiation. The specific results of each gene. The first thing to look at is the result of the change in the peak value in MeRIP-seq. At the same time, the result of the corresponding m6A expression level and the result of the protein expression relative to the protein expression of different groups will also be looked at
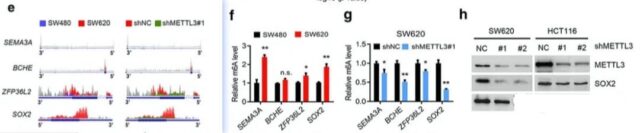
METTL3 promotes the stemness of colon cancer cells in vitro and the migration ability of tumor cells
Since the function of METL3 has been determined through the above analysis, the author’s next step is to do basic experiments. The purpose of the basic experiments is to determine how METL3 plays the role of cell stemness and tumor migration ability. Specific experiments are also regular things. If you are interested, you can take a look at it yourself. Due to the length of the content of the official account, we will not introduce the specific results.
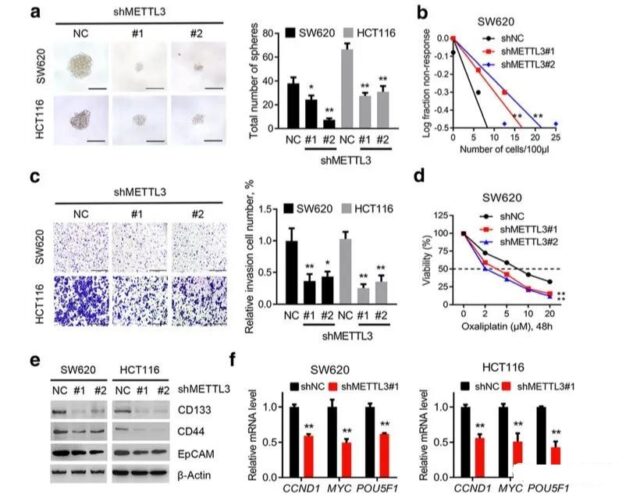
IGF2BP2 can improve the stability of SOX2mRNA through m6A methylation
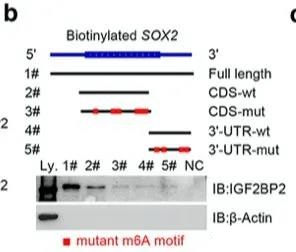
Since the status of SOX2 m6A requires additional genes to maintain, the author did an RNA pull down experiment to find genes related to SOX2 and m6A at the same time, and finally found that IGF2BP2 can bind to SOX2 RNA.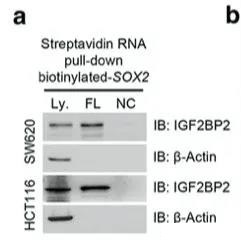
For the prediction of its combined position, the author uses the m6A-related database we introduced earlier, which is also the earliest m6A database: SRAMP.
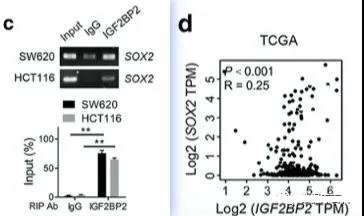
Further, in order to determine its direct binding relationship, the author also did a related RIP experiment. At the same time, the TCGA public database was used to observe the colinearity of SOX2 and IGF2BP2. For this forecast, the GEPIA database is used. Regarding the GEPIA database, we have also introduced the basic usage method. Probably the most famous TCGA expression related database introduction (1) Probably the most famous TCGA expression analysis database (2)
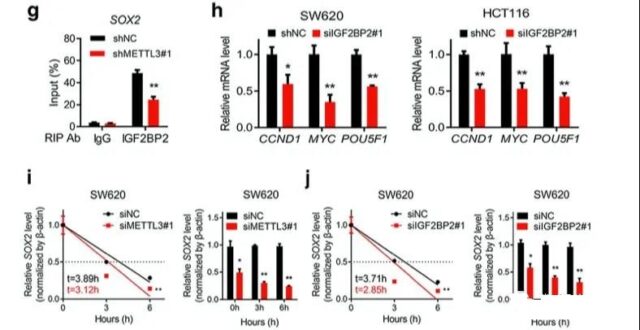
The last is a knockout verification experiment to illustrate the relationship between SOX2, IGF2BP2 and METL2
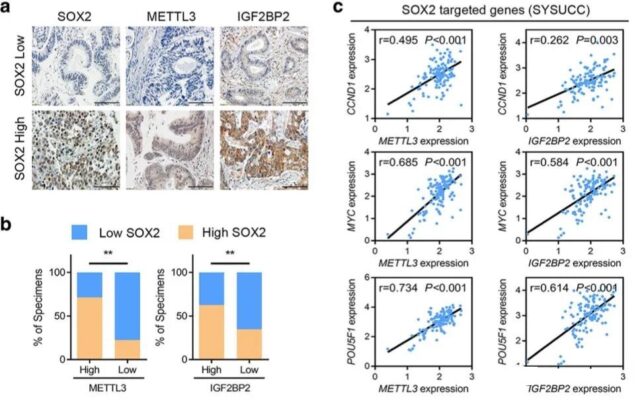
The clinical correlation of METL3, SOX2 and IGF2BP2 in colon cancer
After the basic experiment is finished, in order to enrich the content, we can go to the clinical test book to observe the correlation of these mutually regulated genes finally found, so the author has done relevant experiments and analysis in the clinical data.
(source:internet, reference only)
Disclaimer of medicaltrend.org



