Cyclin-dependent kinase inhibitor CDKN2C is not a host factor for HBV
- Normal Liver Cells Found to Promote Cancer Metastasis to the Liver
- Nearly 80% Complete Remission: Breakthrough in ADC Anti-Tumor Treatment
- Vaccination Against Common Diseases May Prevent Dementia!
- New Alzheimer’s Disease (AD) Diagnosis and Staging Criteria
- Breakthrough in Alzheimer’s Disease: New Nasal Spray Halts Cognitive Decline by Targeting Toxic Protein
- Can the Tap Water at the Paris Olympics be Drunk Directly?
Cyclin-dependent kinase inhibitor CDKN2C is not a host factor for HBV
Cyclin-dependent kinase inhibitor CDKN2C is not a host factor for HBV. From the non-proliferative state to the proliferative state, or vice versa, from the proliferative state to the non-proliferative state, the various omics changes of cells including the expression profile are extremely significant. Among them, the primary human hepatocytes (PHH) in a short period of time after culture in vitro, that is, the transition from the silent state of division to the active proliferation state, so that the expression of a series of hepatocyte-specific genes including NTCP and HNF4a are significantly down-regulated , The performance is no longer sensitive to HBV infection, and the ability to support HBV replication is significantly reduced.
Therefore, in theory, any factor that can affect the cell proliferation state may affect the infection and replication of HBV. These factors do not directly affect the infection and replication of HBV and should not be classified as host factors of HBV.
Recently, in a study published by Carla Eller et al. in Nature Communication (NC for short), they found that cyclin D (CDK4/6) was screened through the whole genome function of Huh7 cells with active proliferation. The suppressor gene cyclin-dependent kinase inhibitor CDKN2C can promote HBV replication, and CDKN2C is called the host factor of HBV. Since in their study, cell cycle arrest is essential for CDKN2C to enhance HBV replication, we wonder whether CDKN2C can affect HBV infection and replication in a non-cell cycle control manner.
In order to reassess the role of CDKN2C in HBV replication in patients with chronic hepatitis B (CHB), we first analyzed the data of 90 CHB patients in the same GSE83148 data set as the NC article, but the CDKN2C expression level was not observed There was a significant difference between patients with HBV DNA load <10⁶ IU/mL and patients ≥ 10⁶ IU/mL (Figure 1A). In order to verify the above results, we analyzed another liver transcriptome data set (GSE84044) containing 124 HBV-related liver fibrosis patients. Similarly, no correlation between liver CDKN2C expression levels and serum HBV gene expression levels was observed. (R = 0.115, P = 0.255) (Figure 1B). Next, we analyzed the third data set GSE94660, which comes from the RNA-seq data of 21 HBV infection-related hepatocellular carcinoma (HCC) patients. The results show that the expression of CDKN2C is related to the HBV in malignant hepatocellular carcinoma tissues. There was also no correlation with RNA levels (r = 0.367, P = 0.107) (Figure 1C).
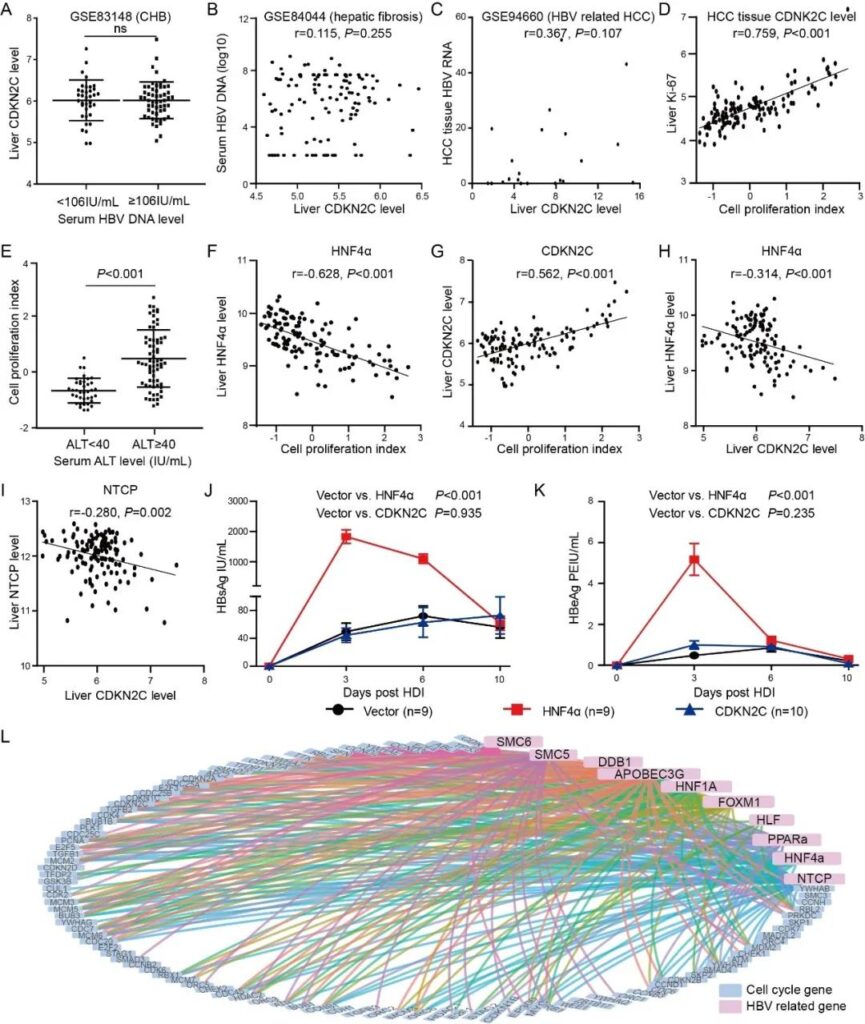
Figure 1: CDKN2C is not a host factor of hepatitis B virus.
- A, In the GSE83148 data set, the expression level of CDKN2C in the liver of CHB patients with serum HBV DNA <10⁶ IU/mL and ≥10⁶ IU/mL.
- B, in the GSE84044 data set, the relationship between the expression of liver CDKN2C and serum HBV DNA in patients with HBV-related liver fibrosis.
- C, the correlation between the expression level of CDKN2C (FPKM) in HCC tissues and the level of liver HBV RNA in the GSE94660 data set.
- D, The correlation between hepatocyte proliferation index and Ki-67 expression in GSE83148 data set.
- E, Compare the hepatocyte proliferation index of chronic hepatitis B patients with different serum ALT levels in the GSE83148 data set.
- F&G, the correlation between hepatocyte proliferation index and the expression of HNF4α and CDKN2C in the GSE83148 data set. H&I, in the GSE83148 data set, the correlation between liver CDKN2C levels and HNF4α and NTCP expression levels.
- J&K, serum HBsAg and HBeAg levels at different time points after hydrodynamic injection into the tail vein of mice.
L, Analysis of the expression correlation between cell cycle-related genes and HBV-dependent host genes. The blue dots represent cell cycle genes, and the pink dots represent HBV-dependent host genes. - The line between the two points indicates that the expression levels of the two genes are highly correlated, with a cutoff value of | r |> 0.4 and fdr <0.05.
According to reports, the expression of CDKN2C is significantly up-regulated during the G1/S transition, and it can block the cell cycle. There are also reports in the literature that the expression level of CDKN2C is related to the terminal differentiation of adipocytes (Morrison and Farmer 1999).
Considering that HBV replication is significantly affected by the growth and proliferation status of infected hepatocytes, the relationship between the expression level of CDKN2C and the proliferation status of hepatocytes in CHB patients is worth exploring. We first selected 92 genes detected by GSE83148 among 116 cell cycle-related genes annotated by KEGG database, and established a cell proliferation index model through factor analysis.
This new index can better reflect the proliferation status of hepatocytes because it is positively correlated with the Ki-67 level in the liver tissues of these CHB patients (r = 0.759, P <0.001) (Figure 1D). Interestingly, the cell proliferation index of patients with higher serum ALT levels was significantly higher than that of patients with normal ALT (UNL, 40 IU/mL) (Figure 1E), indicating that necrotizing inflammatory damage leads to compensatory proliferation of liver cells.
As expected, the cell proliferation index score was significantly negatively correlated with the HBV-dependent host gene HNF4α (r = -6.228, P <0.001) (Figure 1F), which is consistent with previous reports. However, contrary to the report of Eller et al., we found that the expression of CDKN2C was significantly positively correlated with the cell proliferation index in liver tissue specimens derived from CHB (r = 0.562, P <0.001) (Figure 1G). As mentioned above, as an inhibitor of CDK4/6, CDKN2C can prevent the transition of the cell cycle from G1 to S phase.
Based on the fact that the expression of CDKN2C in the liver tissue of CHB patients is positively correlated with the proliferation index, we have reason to infer that the increase in the expression level of CDKN2C is only a secondary response to the transition of hepatic parenchymal cells from resting to rapid proliferation. The “brake” function of the patient’s liver cell proliferation.
Since several studies have reported the inhibitory effect of cell proliferation on HBV replication, we studied the correlation between CDKN2C expression level and a series of known HBV-related host factor gene expression in CHB patients in the GSE83148 dataset.
Eller et al. reported that CDKN2C expression in tumor cells can enhance HBV RNA transcription by up-regulating a series of HBV-related host factor genes known to enhance HBV transcription, such as HNF4α, HLF and PPARα.
In contrast, we found that the mRNA level of CDKN2C is comparable to HNF4α (r = -0.314, P <0.001), NTCP (r = -0.280, P = 0.002) (Figure 1H, 1I), HLF (r = -0.209, P = 0.021) and PPARα (r = -0.317, P <0.001) were negatively correlated (Supplementary Figure S1A and Supplementary Figure S1B).
In addition, similar results were obtained in the data (GSE84044) of patients with liver fibrosis associated with HBV infection (Supplementary Figures S1C–S1F). Since CDKN2C is not positively correlated with the expression of HBV-dependent genes in liver tissues of CHB patients, it is unlikely that CDKN2C can enhance HBV replication by up-regulating these HBV-related host factor genes.
Because most liver cells are in a quiescent (G0) state, we believe that CDKN2C is unlikely to inhibit cell proliferation in vivo and promote HBV replication. To verify this hypothesis, we injected HNF4α or CDKN2C expression plasmids and the same amount of 1.2mer HBV plasmids into mice by hydrodynamic injection (HI) into the mouse tail vein, and used the vector pcDNA3.1 as a control.
The mouse serum was collected at 3, 6 and 10 days after HI, and the levels of HBeAg and HBsAg in the serum were measured. As expected, the HBeAg and HBsAg levels of the HNF4α group were more than 15 times higher than that of the control group, but there was no significant difference between the CDKN2C group and the control group (Figure 1J, 1K).
The above results indicate that CDKN2C cannot promote HBV replication in vivo. Since then, we used the data in the GSE83148 dataset to further study other known cell cycle-related genes and HBV-dependent host genes (NTCP, HNF4α, HLF, PPARα, HNF-1, HNF-3, APOBEC3, DDB1, SMC5 and SMC6) The relationship between.
We found that almost all 92 cell cycle genes are associated with at least one known HBV-dependent gene (Figure 1L), which indicates that almost all of these cell cycle regulatory genes may affect the expression of HBV-dependent genes, thereby affecting HBV replication.
All in all, we proved here that the increase in CDKN2C expression is positively correlated with the proliferation status of parenchymal liver cells. Since there is no CDKN2C expression in the dominant hepatocytes in the resting state, there is no correlation between CDKN2C expression and HBV replication in HBV infected patients. Animal experiments also confirmed that CDKN2C cannot promote HBV replication in vivo. It is inappropriate to call it HBV host factor.
(source:internet, reference only)
Disclaimer of medicaltrend.org



