Membrane-permeable and highly selective Pin1 covalent inhibitor
- Normal Liver Cells Found to Promote Cancer Metastasis to the Liver
- Nearly 80% Complete Remission: Breakthrough in ADC Anti-Tumor Treatment
- Vaccination Against Common Diseases May Prevent Dementia!
- New Alzheimer’s Disease (AD) Diagnosis and Staging Criteria
- Breakthrough in Alzheimer’s Disease: New Nasal Spray Halts Cognitive Decline by Targeting Toxic Protein
- Can the Tap Water at the Paris Olympics be Drunk Directly?
Membrane-permeable and highly selective Pin1 covalent inhibitor
Membrane-permeable and highly selective Pin1 covalent inhibitor. Protein phosphorylation and the cascade after phosphorylation regulate a variety of cancer-related signal transduction pathways in cells. In cell cycle control, the serine/threonine-proline motif is the only phosphorylation site of many key protein kinases.
It has two completely different cis/trans isomers, and the peptidylproline cis Peptidyl-prolyl cis/trans isomerase NIMA-interacting 1 is the only enzyme found to catalyze the mutual conversion of the cis/trans isomers. This uniqueness determines its irreplaceable role in cell cycle regulation, transcription factor regulation, cell proliferation and differentiation regulation.
Pin1 is closely related to the occurrence and development of many human tumors. It activates at least 43 oncoproteins, inactivates more than 20 tumor suppressors, and down-regulates overall microRNAs. It is the “master” post-phosphorylation regulator of oncogenic signaling networks , Is called the oncogenic signal of tumor development. In addition, studies have shown that Pin1 is necessary for tumorigenesis induced by Ras activation. Therefore, the development of Pin1 inhibitors is of great significance for treating Ras-driven tumorigenesis and circumventing Ras’ difficult targets.
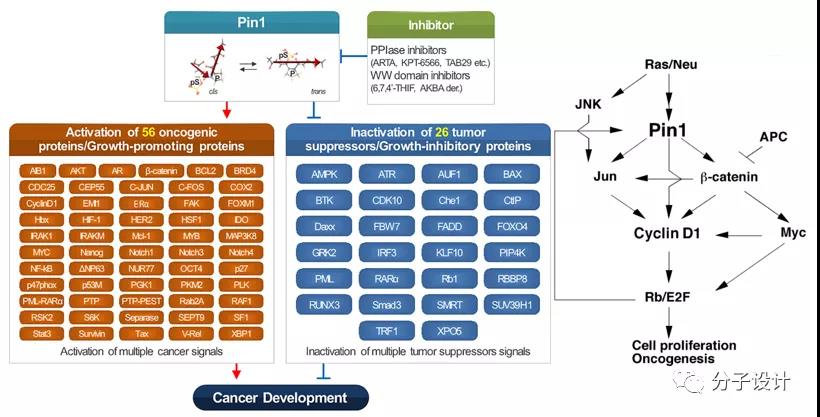
However, the development of Pin1 inhibitors is quite challenging. The realization of the inhibitory effect requires the anionic compound to interact with the phosphate binding pocket of Pin1, which will sacrifice the cell permeability of the compound, and when the membrane permeability is emphasized, the compound may lose the corresponding selectivity.
The previously reported compounds are not immune to this problem. Researchers at Harvard Medical School in the United States have recently developed a Pin1 covalent inhibitor BJP-06-005-3 that combines selectivity and membrane permeability, which can be used to study the function and mechanism of Pin1 in tumorigenesis and development. .
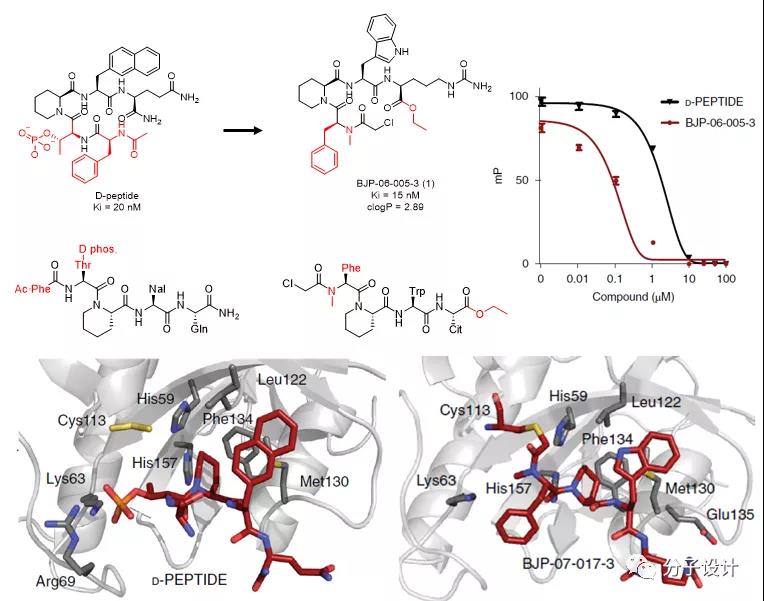
This work is based on the previously reported Pin1 non-natural peptide inhibitor D-PEPTIDE, which has high affinity for protein (Ki = 20 nM), but cannot penetrate the membrane into cells. Based on a rational design, the researchers replaced threonine forming the phosphate with phenylalanine, replaced the amino end at the carbon end with ethyl ester, and methylated the amide at the nitrogen end to eliminate hydrogen bond donors, thereby improving peptide inhibitors The cell permeability.
Later, a covalent binding strategy is considered to compensate for the loss of affinity caused by the replacement of cell permeability groups. With reference to the eutectic structure of D-PEPTIDE and the identification of cysteine reactivity at the omics level, α-chloroacetamide was introduced as an electrophilic group to form a covalent coupling with Cys113 at the nitrogen end of Pin1, thereby obtaining a high balance Affinity and high permeability molecule BJP-06-005-3 (Ki = 15 nM, clogP = 2.89).
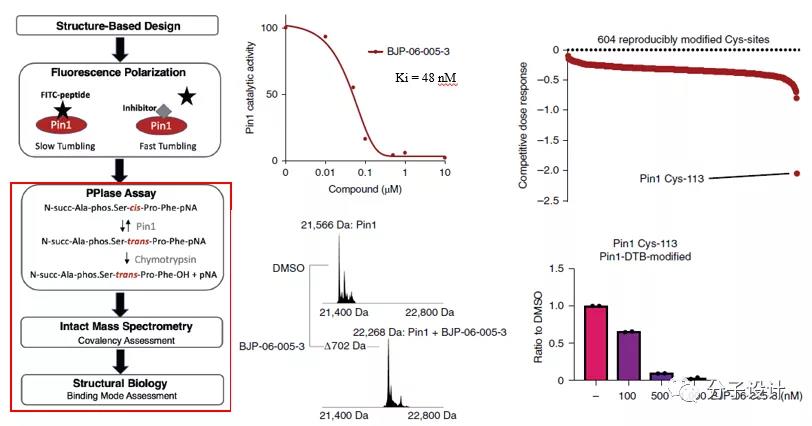
After that, the researchers evaluated the effectiveness, covalency and selectivity of BJP-06-005-3. The ability of BJP-06-005-3 to inhibit the enzyme activity of Pin1 was tested by the PPIase experiment coupled with chymotrypsin, and its apparent Ki value was 48 nM. With the aid of mass spectrometry and structural biology, it was proved that BJP-06-005-3 showed 100% covalent labeling to Pin1 and the modification site was indeed Cys113.
Most importantly, the researchers confirmed that BJP-06-005-3 is unique in the covalent inhibitor target site identification (CITe-Id) experiment with the help of dethiobiotin (DTB) coupled probe (BJP-DTB) Selectivity-In the entire proteome labeling sites of HEK293 lysate, Cys113 of Pin1 is the only cysteine that shows a dose-dependent binding to BJP-06-005-3.
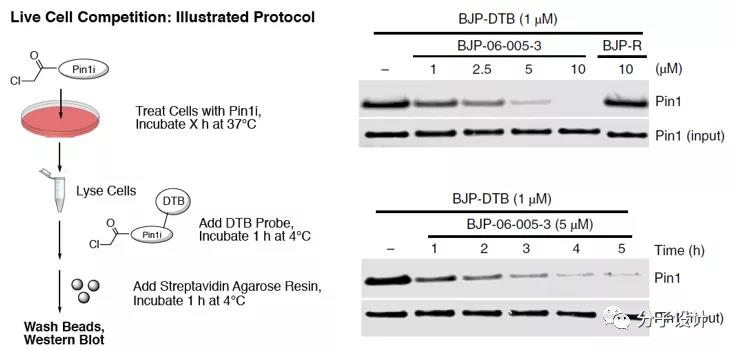
The cell permeability of BJP-06-005-3 was also evaluated in a live cell competition experiment, which showed a dose-dependent and time-dependent competition for the binding of BJP-DTB to Pin1.
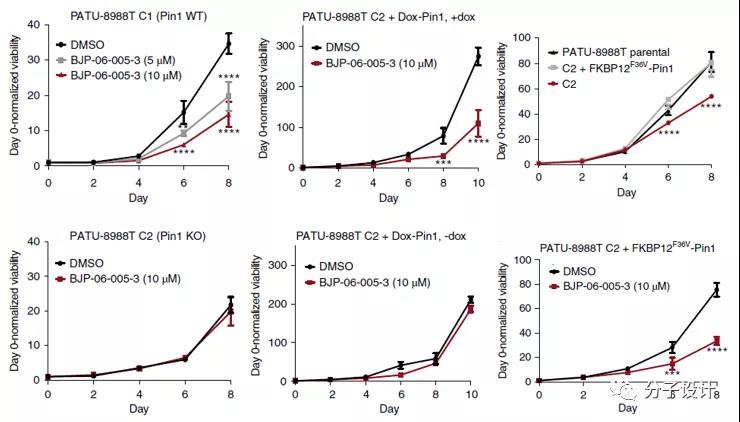
The researchers also identified BJP-06-005-3 mediated the function of Pin1 on tumor cells in the PDAC model. BJP-06-005-3 can reduce the transformation ability of pancreatic cancer cell line (PATU-8988T C1) in a dose-dependent and time-dependent manner. In Pin1 knockout cell line (PATU-8988T C2), whether it is Amycin The expression of Pin1 induced by Dox or the transfer of Pin1 fusion protein (FKBP12F36V-Pin1) can also effectively restore the inhibition of BJP-06-005-3 on tumor cell viability.
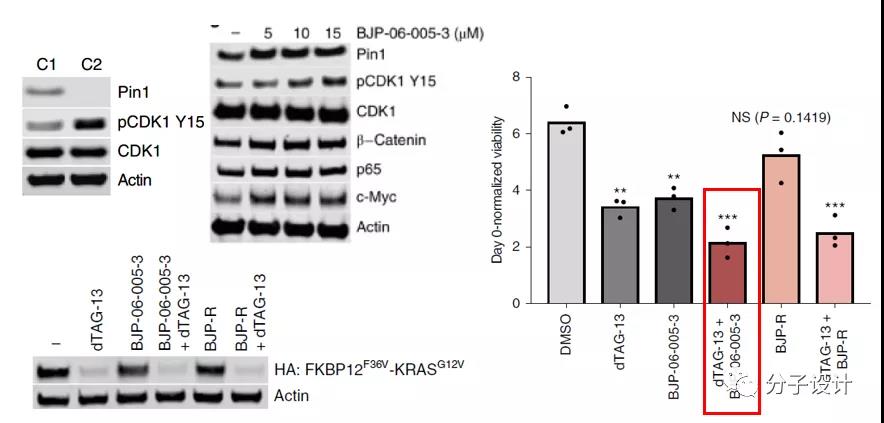
Considering the important function of Pin1 in cell cycle regulation, the researchers also explored the effect of BJP-06-005-3 on cell cycle progression. BJP-06-005-3 can increase the phosphorylation of tyrosine 15 of CDK1, a key factor of mitosis, in a dose-dependent manner, thereby moderately blocking the cell cycle.
In addition, Pin1 is an essential factor for Ras to drive tumorigenesis. Further exploration found that Pin1 inhibition can cooperate with KRAS mutation degradation to play an anti-tumor cell proliferation effect.
In this study, under the guidance of rational design, through repeated optimization of the potency, selectivity and membrane permeability of peptide inhibitors, the current Pin1 membrane-permeable inhibitor BJP-06-005-3 with the best selectivity was obtained. BJP-06-005-3 can cooperate with the degradation of KRASG12V to play an anti-tumor cell proliferation function.
More importantly, BJP-06-005-3 is a powerful multifunctional tool compound that can be used to further explore the role of Pin1 in Ras-driven tumorigenesis and development, thereby providing new ideas for difficult-to-treat tumors such as pancreatic cancer. The solution ideas.
(source:internet, reference only)
Disclaimer of medicaltrend.org



