COVID-19 and its vaccines: Comprehensive Interpretation
- Normal Liver Cells Found to Promote Cancer Metastasis to the Liver
- Nearly 80% Complete Remission: Breakthrough in ADC Anti-Tumor Treatment
- Vaccination Against Common Diseases May Prevent Dementia!
- New Alzheimer’s Disease (AD) Diagnosis and Staging Criteria
- Breakthrough in Alzheimer’s Disease: New Nasal Spray Halts Cognitive Decline by Targeting Toxic Protein
- Can the Tap Water at the Paris Olympics be Drunk Directly?
COVID-19 and its vaccines: Comprehensive Interpretation
- Should China be held legally responsible for the US’s $18 trillion COVID losses?
- CT Radiation Exposure Linked to Blood Cancer in Children and Adolescents
- Can people with high blood pressure eat peanuts?
- What is the difference between dopamine and dobutamine?
- What is the difference between Atorvastatin and Rosuvastatin?
- How long can the patient live after heart stent surgery?
COVID-19 and its vaccines: Comprehensive Interpretation.
This article explains in detail the components of COVID-19, the pathogenic mechanism and the corresponding vaccine development.
What is COVID-19 and SARS-CoV-2
With the outbreak of the COVID-19 epidemic and the continuous progress of scientific research, the two English words COVID-19 and SARS-CoV-2 have been constantly appearing in people’s vision.
What is this and what are they related to? COVID-19 is the name of the disease caused by the COVID-19 virus named by the WHO.
COVID is a combination of the three words Corona (corona), Virus (virus) and Disease (disease), and 19 represents 2019. SARS-CoV-2 is the name of the virus that causes COVID-19.
The R0 value of SARS-CoV-2
SARS-CoV-2 is extremely infectious. Since the outbreak of 2019.11, it has spread to 188 countries and 25 territories.
R0 is a parameter used to measure infectiousness, and its size depends on various biological, environmental and social behavioral factors.
The R0 value of SARS-CoV-2 is 2-3, while the R0 value of Spanish influenza is 0.9-2.1.
SARS-CoV-2, SARS-CoV and MERS-CoV
Coronavirus is a +RNA enveloped virus whose surface is mainly covered by S protein. Genetic analysis showed that SARS-CoV-2 and SARS-CoV are 79% similar.
SARS-CoV-2 and SARS-CoV share a common host receptor, human angiotensin-converting enzyme 2 (hACE2), but SARS-CoV-2 has a stronger ability to bind hACE2 than SARS-CoV.
Although the two are very similar, they have no cross-protection effect. The similarity between MERS-CoV and SARS-CoV-2 is 50%, and the receptor for MERS-CoV is DPP4 instead of hACE2.
Non-structural and structural proteins in SARS-CoV-2
The SARS-CoV-2 genome contains at least 10 ORFs. ORF1ab is converted into a polyprotein, which is processed into 16 non-structural proteins (NSP).
These NSPs have multiple functions, such as genome replication, induction of host mRNA cleavage, membrane rearrangement, autophagosome production, NSP polyprotein cleavage, capping, tailing, methylation, RNA double-strand unwinding, etc. Play an important role in the life cycle.
In addition, SARS-CoV-2 contains 4 structural proteins, namely spike (S), nucleocapsid (N), envelope (E) and membrane (M), all of which are encoded by the 3’end of the viral genome.
Among the four structural proteins, S protein is a large multifunctional transmembrane protein, which plays an important role in the process of virus adsorption, fusion, and injection into host cells.
1) The S protein is composed of S1 and S2 subunits, and each subunit can be further divided into different functional domains. The S1 subunit has two domains: NTD and RBD, and RBD contains conservative RBM. The S2 subunit has three domains: FP, HR1 and HR2. The S1 subunit is arranged at the top of the S2 subunit to form an immunodominant S protein.
The virus uses the host transmembrane protease Serine 2 (TMPRSS2) and the endosomal cysteine protease CatB/L to enter the cell.
TMPRSS2 is responsible for the cleavage of the S protein to expose the FP region of the S2 subunit, which is responsible for initiating endosome-mediated host cell entry.
It shows that TMPRSS2 is a host factor necessary for virus entry. Therefore, the use of drugs that inhibit this protease can achieve therapeutic purposes.
SARS-CoV-2 targets cells by binding to human hACE2 via S protein. Coronavirus uses conformational masking and sugar masking to make S protein escape the immune system. Frozen EM revealed that the S protein extradomain trimer has two conformations: open and closed. This conformational change is crucial for binding to hACE2, because the opening of the trimer can expose RBM.
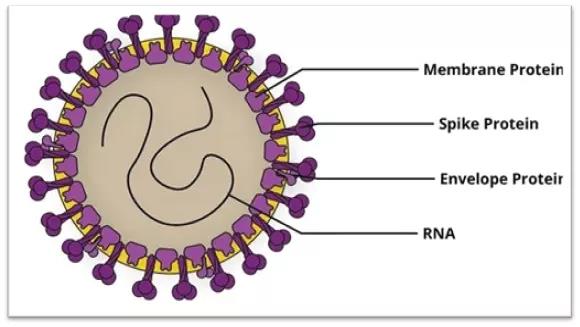 Figure 1 TMPRSS2 inhibitor nafamostat has therapeutic effects
Figure 1 TMPRSS2 inhibitor nafamostat has therapeutic effects
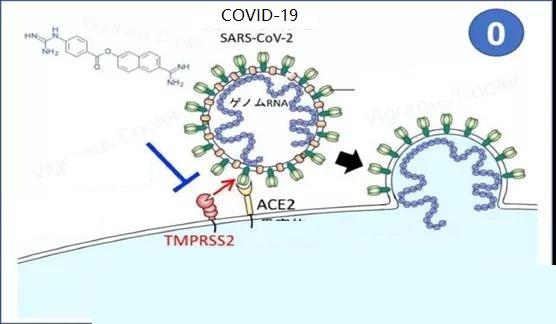 Figure 2 Schematic diagram of SARS-CoV-2 structural protein
Figure 2 Schematic diagram of SARS-CoV-2 structural protein
2) The E protein forms the E channel, which plays a variety of functions in the virus replication cycle, including integration, release, and pathogenic mechanisms.
These ion channels exist in the form of homopentamers, each subunit containing 50-120 amino acids.
The E channel contains at least one TMD, which facilitates the connection with the host cell membrane.
3) In the virus particle, the M protein is the central organizer, and plays a vital role in the morphogenesis and assembly of SARS-CoV-2 by interacting with essential structural proteins.
The combination of M and N proteins stabilizes the N protein and RNA complex, as well as the inner core of the virus.
In SARS-CoV, M protein has also been shown to induce host cell apoptosis. The N protein has the effect of stabilizing the ssRNA of virus particles, is an anti-viral RNAi antagonist, and is also responsible for inhibiting host cells from entering the S phase (DNA synthesis phase).
Vaccine development requires careful verification and adverse reaction assessment.
The target population of the vaccine includes high-risk groups over 60 years old (especially people with chronic diseases), front-line medical workers and people in the essentials industry.
S protein is the main antigen for preparing vaccines, and the neutralized serum can provide protection for the body.
The SARS-CoV-2 full-length 5500 genome has been released at NCBI, providing a basis for vaccine development.
As of July 2, 2020, there are 158 SARS-CoV-2 vaccine candidates worldwide, of which 135 are in the preclinical or research and development stage.
The vaccine development platform includes
- Protein subunit vaccines,
- Viral vector vaccines,
- Nucleic acid vaccines (DNA, RNA),
- Live attenuated vaccines
- Passive immunization.
Each vaccine has its own advantages and disadvantages.
1. Protein subunit vaccine
Such vaccines are usually low in immunogenicity and need to be used with adjuvants to induce immunity.
The adjuvant enhances the biological half-life of the antigen material or improves the immunomodulatory cytokine, and makes up for the defects of the subunit vaccine.
SARS-CoV-2S protein is the most suitable antigen for inducing neutralizing antibodies. The virus enters cells through endocytosis mediated by S protein and hACE2 receptor.
S glycoprotein has conformational changes, so in order to better induce antibody response, its antigen structure and conformation should be retained.
1) NVX-CoV2373
NVX-CoV2373 is a recombinant nano-vaccine based on the pre-fusion S protein. The protein is stably expressed by baculovirus, and is designed to use Matrix-M adjuvant to enhance the immune effect.
In animal experiments, a single immunization can obtain high levels of antibodies, which can block the binding of SARS-CoV-2 to hACE2 receptors.
2) PittCoVacc
A microinjection formulation of rSARS-CoV-2 S1 and rSARS-CoV-2-S1fRS09 components. In preclinical experimental mouse models, statistically significant antibodies can be detected 2 weeks after vaccination.
Moreover, the immunogenicity of the vaccine remains after sterile treatment with gamma rays. Before the booster immunization, a statistically significant level of antibodies can be obtained to make the vaccine feasible.
3) Three-component antigen vaccine
A multi-antigen virus-like particle co-expressing SARS-CoV-2 recombinant S, M, E by the Saccharomyces cerevisiae expression platform (D-CryptTM), which is expected to enter preclinical trials.
2. Viral vector vaccine
Viral vector vaccines can provide long-term high-level expression of antigen proteins, induce CTLs, and ultimately eliminate viral infections.
1) Ad5-nCov
A vaccine of SARS-CoV-2 recombinant spike protein expressed by recombinant, replication-deficient type 5 adenovirus (Ad5) vector.
The optimized full-length S protein gene and plasminogen activation signal peptide gene were loaded into the E1 and E3 deleted Ad5 vector. The vaccine is constructed by the Admax system derived from Microbix Biosystem.
In phase I clinical trials, RBD (S1 subunit receptor binding domain) and S protein neutralizing antibody increased by 4 times 14 days after immunization, reaching a peak at 28 days. CD4+T and CD8+T cells reached a peak 14 days after immunization.
The existing Ad5 immune resistance partially limits the antibody and T cell response. This study will be further conducted in the 18-60 age group, receiving 1/3 of the study dose, and follow-up for 3-6 months after immunization.
2) Coroflu
M2SR is a self-limited influenza virus, into which the SARS-CoV-2 spike protein gene sequence is inserted to prepare a vaccine.
The vaccine can also express influenza virus hemagglutinin and induce a double immune response.
M2SR cannot replicate due to lack of M2 gene, but it can enter cells and induce immune response.
It can be immunized through the nose (imitating the natural infection mode) to activate a variety of immune system modes.
Compared with intramuscular injection, a high level of immune effect can be obtained.
3) LV-SMENP-DC
Use SMENP minigenes to infect engineered dendritic cells with a lentivirus expressing SARS-CoV-2 structural protein and protease conserved domains to obtain LV-SMENP-DC vaccine.
Subcutaneous immunity activates CTLs.
4) ChAdOx1
The codon-optimized SARS-CoV-2 S protein coding sequence was introduced into the 5’end of the tissue plasminogen activation (tPA) leader sequence and cloned into the shuttle plasmid.
The shuttle plasmid encodes the main early genes of human cytomegalovirus (IECMV) between the Gateway recombination cloning sites, as well as the tetracycline manipulation (TetO) site and bovine growth hormone (BGH) polyadenylation signal.
The SARS-CoV-2S gene was inserted into the E1 site of the ChAdOx1 adenovirus genome.
The virus was propagated in the T-Rex293HEK (human embryonic kidney 293) cell line and purified by CsCl gradient ultracentrifugation.
In preclinical trials, animal experiments with intramuscular inoculation showed good immune responses. The vaccine has entered Phase II clinical trials, increasing the number of samples for evaluation.
3. mRNA vaccine
The mRNA vaccine is not infectious and there is no risk of insertion mutation. Including non-replicating RNA and virus-driven self-replicating RNA.
The optimization can enhance the stability of the mRNA vaccine, reduce the immunogenicity of the carrier, avoid anti-carrier immunity, and achieve repeatable immunization.
This platform enhances the ability of rapid vaccine development, has the advantages of design flexibility and imitating the natural infection process.
1)mRNA-1273
The mRNA encoding the full length of SARS-CoV-2 and the fusion pre-spike protein is encapsulated into lipid nanoparticles to form mRNA-1273 vaccine.
It can induce a high level of S protein specific antiviral response. It can also consist of inactivated antigens or subunit antigens.
The vaccine was quickly approved by the FDA and has entered phase II clinical trials.
The company has published the antibody data of 8 subjects who received different immunization doses. The 25ug dose group achieved a similar effect to the antibody level during the recovery period.
The 100ug dose group exceeded the antibody level during the recovery period. The vaccine was basically safe and tolerable in the 25ug and 100ug dose groups, while the 250ug dose group had 3 grade system symptoms.
2) BNT162b1
BNT162b1 is a codon-optimized RBD trimer mRNA vaccine encoding SARS-CoV-2.
The addition of T4 fibrin derived folded trimer structure to the RBD antigen enhances the immunogenicity of the vaccine. The mRNA is encapsulated in 80nm ionizable cationic lipid nanoparticles, which can ensure its effective delivery.
Phase 1/2 clinical trials revealed that the level of RBD-specific IgG antibodies was 8-46.3 times higher than the geometric mean concentration of serum during the rehabilitation period.
The geometric mean concentration of SARS-CoV-2 neutralizing antibody is only 1.8-2.8 times the serum level during the recovery period.
Moderately short-lived local and systemic reactions did not appear as side effects. The immune response and safety after 2 weeks of the second immunization have not been evaluated.
4. DNA vaccine
The introduction of antigen-encoding DNA and adjuvants as vaccines is the most innovative vaccine method. The transfected cells stably express the transgenic protein, similar to live viruses.
The antigen will be endocytosed by immature DC, and finally provide antigen to CD4+T and CD8+T cells (differentiated by MHC) to induce humoral and cellular immunity.
INO-4800
Use the SARS-CoV-2 codon to optimize the S protein sequence and attach the IgE leader sequence. The SARS-CoV-2 IgE spike sequence is synthesized and digested, loaded into the pGX0001 expression plasmid, and driven by IE CMV and BGH polyadenylation signals.
An effective immune response can be induced 7 days after immunization. It has entered phase I clinical trials and is estimated to be completed before July.
Subjects received 1.0 mg of INO-4800 through electroporation. Both intradermal injection and electrotransformation in healthy subjects showed safety and tolerance.
5. Live attenuated vaccine
DelNS1-SARS-CoV2-RBD
Basic influenza vaccine, delete NS1 gene. Expression of SARS-CoV-2 RBD domain. Cultured in CEF and MDCK (canine kidney cells) cells. It is more immunogenic than wild-type influenza virus and can be administered by nasal spray.
6. Others
With the deepening of the understanding of the structure and genome of SARS-CoV-2, various vaccines have been developed rapidly.
It has potential immune effects and side effects. Vaccine development is a long-term and arduous process that requires long-term evaluation. Various biological companies use different technologies to develop vaccines.
BAT uses new, fast-growing tobacco to prepare the COVID-19 vaccine. Tianjin University developed an oral vaccine through S. cerevisiae carrying S protein.
The immune system is activated by heat shock protein (HSP), a fusion protein is formed between HSP and avidin, and vaccine preparation is realized in the form of biotinylated immune peptide.
7. Passive immunity
It is the use of antibodies to treat diseases, including the serum of convalescent patients, polyclonal antibodies produced in other animals, and neutralizing monoclonal antibodies or humanized antibodies produced by hybridoma technology.
1) Convalescent serum
As of now, there is no specific treatment for COVID-19. Convalescent serum (CP) therapy is unanimously considered as an emergency response to disease outbreak.
It has been used to treat and prevent SARS, MERS, H1N1, measles, mumps, etc.
One possible explanation is that neutralizing antibodies can overcome viremia, block new infections, and accelerate the clearance of infected cells.
In order to evaluate the effect of CP, various experiments have been carried out, and it has been consistently shown that neutralizing antibodies can improve the clinical condition of critically ill patients without death.
CP treatment dose is not specified, and randomized clinical trials are needed to exclude the effects of other drugs on the one hand and evaluate the effect and safety of CP therapy.
Critically ill patients with similar symptoms, such as high blood pressure, cardiovascular disease, thoracic disease, chronic renal failure, etc. are also included in the study.
They all enter the ICU and receive ventilators.
All these patients are treated with antiviral, antibacterial or antifungal drugs.
Compared with the control, the CP treatment group had no significant differences in baseline characteristics, but showed great differences in clinical results.
Such as body temperature normalization, lung disease absorption, ARDS acute respiratory distress syndrome alleviation, remove the ventilator, reduce mortality.
The viral load is 0 after 7-37 days of CP treatment. Inflammatory factors such as CRP, procalcitonin, and IL-6 decrease, while RBD-specific IgM and IgG increase.
These uncontrolled, non-randomized trials hinder the evaluation of the therapy by scientific researchers and need to continue to be evaluated in clinical trials.
2) Monoclonal antibodies
Monoclonal antibodies are prepared in the laboratory, derived from one patient, and have a monovalent affinity.
Monoclonal antibodies do not have the disadvantages of convalescent plasma in terms of specificity, safety, low risk of blood-borne infection, purity, and other factors.
A variety of monoclonal antibodies have been developed for anti-tumor, anti-platelet or anti-viral therapy.
The SARS-CoV specific human monoclonal antibody CR3022 can be combined with RBD, indicating that it has a therapeutic effect and can be used alone or in combination.
In order to ensure the efficacy, various monoclonal antibodies can be combined for treatment to identify different virus targets, effectively treat and block virus escape.
There are currently more than 61 patents for preparation of SARS\MERS\diagnostic antibodies.
Other 38 patents develop antibodies that target host proteins such as IL-6/IL-6R, TLR3, CD16, ITAM, DC-SIGN, ICAM-3, and IP-10/CXCL10. These antibodies can resist the cytokine storm and alleviate SARS-CoV-2 infection.
Tocilizumab, IL-6 receptor antibody can control hyperinflammatory lung symptoms. Tocilizumab can block the cytokine IL6, thus inhibiting the inflammatory cascade.
Clinical trials are needed to further determine the effectiveness of monoclonal antibodies.
Difficulties in the development of SARS-CoV-2 vaccine
1) Vaccine development takes a long time, and disease outbreaks are rapid
The SARS-CoV-2 epidemic has led to a substantial end to economic activities. The prevention and treatment of SARS-CoV-2 requires global scientists to work together to establish innovative partnerships with pharmaceutical giants and medical start-ups to re-use drugs, develop vaccines and equipment , Hindering the progress of this overwhelming epidemic.
The vaccine development process is an arduous process, including preclinical and phase III clinical stages. In order to accelerate vaccine research and development, with sufficient data support, consider skipping certain stages and speed up vaccine production through rapid regulatory review, approval, manufacturing and quality control.
Therefore, the emergence of the new coronavirus has forced the scientific community to use unconventional methods to accelerate the development of vaccines.
But no matter when, vaccine development should have the following principles: high efficiency, only minor or short-term adverse effects, and will not cause serious diseases.
The vaccine must be suitable for people of all ages, including pregnant and lactating women. A single dose of immunization can provide rapid protection with at least one year of protection.
2) Conventional animal models are not suitable for SARS-CoV-2 vaccine research
Vaccine development and testing require support from animal models, which must show disease processes similar to humans.
However, due to the difference between human and mouse hACE2 receptors, standard inbred mice are not susceptible to SARS-CoV-2 infection, which requires the development of transgenic mice that express hACE2 receptors.
In order to study the hACE2 transgenic mouse model and primate macaque model developed by SARS-CoV, it is currently necessary to solve the problem of stable reproduction and adaptation to global use to meet research needs.
The SARS-CoV-2 virus isolate can effectively replicate in the lungs of Syrian hamsters, and the lungs of infected hamsters show pathological changes similar to those of patients with COVID-19 pneumonia. In addition, the immune response produced by the infected hamster can resist re-infection.
Infusion of convalescent serum into healthy hamsters can prevent the virus from replicating in the lungs.
These experiments show that Syrian hamsters can be used as animal models for studying the pathogenesis of SARS-CoV-2 and evaluating antiviral drugs and immunotherapy.
However, the assessment of vaccine dependence on immune enhancement cannot be inferred from animal models and needs to be confirmed in phase III clinical trials.
3) Antibody-dependent enhancement (ADE) and viral genome mutation
In 1964, the Hawkes team of the Australian National University first proposed the “antibody-dependent enhancement” (ADE) hypothesis in arboviruses.
That is, after some virus-specific antibodies (usually non-neutralizing antibodies) bind to the virus, not only will they not inhibit the virus infection, but will act as a “bridge” to mediate the virus into the cell, thereby enhancing the virus infection Sexual process.
Subsequent studies found that the ADE effect is widely present in a variety of viruses such as dengue fever and other flaviviruses, coronaviruses, HIV, Ebola, etc.
It is a major research focus and problem in epidemiology and vaccine antibody development.
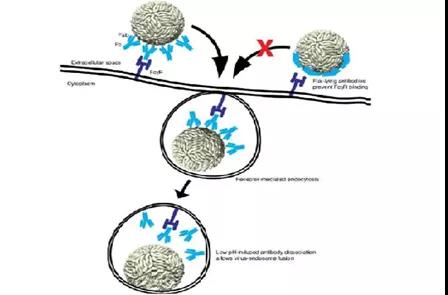 Figure 3 Schematic diagram of antibody ADE
Figure 3 Schematic diagram of antibody ADE
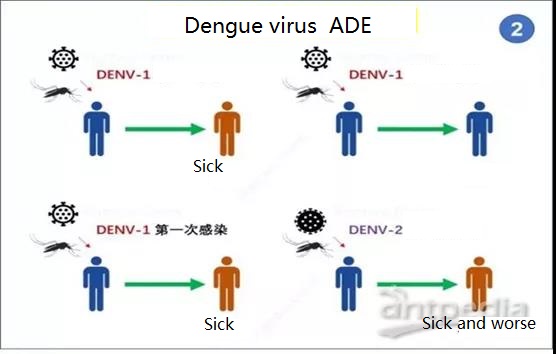
Take dengue virus as an example. Dengue virus has 4 different serotypes (DENV1-4).
When the virus subtype DENV-1 invades the body, the body’s immune cells will produce corresponding anti-DENV-1 neutralizing antibodies.
However, when another serotype virus DENV-2 is re-infected, the previously produced anti-DENV-1 neutralizing antibody will first bind to the DENV-2 virus, but will not neutralize the virus, but will be affected by lgGFc on immune cells.
Body binding promotes the process of virus infecting these cells, thereby aggravating the infection.
Previous studies have found that relying on the ADE effect mediated by Fc receptor (FcR), the virus-antibody complex binds to cells with FcR (including macrophages, monocytes) mainly through the interaction of antibody Fc fragments and cell surface FcR. Cells, B cells, neutrophils and granulocytes, etc.) make the virus adhere to the cell surface and promote infection.
In the ADE effect mediated by the complement receptor (CR), the virus surface protein can bind antibodies of different serotypes to activate the classical pathway of the complement system.
Complement C1q binds to the protein attached to the surface of the virus, thereby bringing the virus and the cell closer to each other by binding to the C1q receptor present on the cell surface, causing the virus to infect the cell.
In addition to the above two, various ADE effect mechanisms such as the ADE effect mediated by virus surface proteins and the ADE effect of antibody mimicking the binding of cell receptors to the virus have also been discovered.
Although there is no obvious ADE effect in the research of new coronavirus antibodies and vaccines, the ADE effect has been found in previous studies of SARS-CoV and MERS-CoV.
Due to the impact of the ADE effect, some vaccines have also caused serious public health events due to the lack of understanding of the ADE effect.
Therefore, in-depth study of the ADE effect mechanism and finding the rules that produce the ADE effect can help us successfully avoid the harm caused by the ADE effect.
At present, it is generally believed that measures such as designing a vaccine that can ensure the induction of high-titer neutralizing antibodies in the body, and can induce both cellular and humoral immunity, can effectively avoid the ADE effect.
Some studies have also found that in HIV vaccine design, if the infection enhancing epitope is removed and the neutralizing epitope is retained, better immune protection can be produced.
In addition, using appropriate drugs to block FcR, combined with vaccines or antibodies to treat viral infections is also an effective strategy to inhibit ADE.
In the development of new coronavirus antibodies, in order to avoid the ADE effect, Shi Rui et al. also introduced LALA mutations in the Fc part of antibody CB6 (CB6-LALA) to reduce the risk of Fc-mediated acute lung injury.
However, in order to successfully avoid the ADE effect, one of the first questions that should be solved is which antibodies and viruses can cause the ADE effect?
What do they have in common?
Only by understanding the specific mechanism and formation rules of the ADE effect can the ADE effect be effectively avoided.
The viral genome is susceptible to mutations, antigen transfer and drift can occur, and spread among the population. Mutations can vary depending on the environmental conditions and population density of the geographic area.
After screening 7,500 samples of infected patients, scientists identified 198 mutations, indicating the evolution of the virus in the human host.
These mutations may form different virus subtypes, which means that even after vaccine immunization, viral infection may occur.
Reference:
1.SimranPreet Kaur, Vandana Gupta.COVID-19 Vaccine: A comprehensive status report.VirusResearch 288 (2020) 198114.https://doi.org/10.1016/j.virusres.2020.198114
2.https://ibook.antpedia.com/x/495814.html
3.https://m.internet.com/a/389888375_120308285/pvid=000115_3w_a&scm=1002.44003c.fe017c.PC_ARTICLE_REC
COVID-19 and its vaccines: Comprehensive Interpretation
(source:internet, reference only)
Disclaimer of medicaltrend.org
Important Note: The information provided is for informational purposes only and should not be considered as medical advice.



