PI3K-AKT pathway
- Normal Liver Cells Found to Promote Cancer Metastasis to the Liver
- Nearly 80% Complete Remission: Breakthrough in ADC Anti-Tumor Treatment
- Vaccination Against Common Diseases May Prevent Dementia!
- New Alzheimer’s Disease (AD) Diagnosis and Staging Criteria
- Breakthrough in Alzheimer’s Disease: New Nasal Spray Halts Cognitive Decline by Targeting Toxic Protein
- Can the Tap Water at the Paris Olympics be Drunk Directly?
PI3K-AKT pathway
PI3K-AKT pathway. In the previous sections, we introduced RTKs (Phase 1 and Phase 2) and MAPK pathway (Phase 3). Today we introduce another important pathway downstream of RTKs and parallel to MAPK—PI3K pathway.
01 The composition and activation method of PI3K
Phosphatidylinositide 3-kinases (PI3K) is an intracellular phosphatidylinositol kinase that has phosphatidylinositol kinase activity as well as serine/threonine (Ser/Thr) kinase activity . PI3K can be divided into 3 categories, of which the most widely studied is type I PI3K. This type of PI3K is a heterodimer consisting of a regulatory subunit p85 (encoded by the PIK3R1 gene) and a catalytic subunit p110 (encoded by the PIK3CA gene) )composition. There are four types of catalytic subunits: α, β, δ, and γ. Among them, α, β, and δ correspond to p85α, p85β or p55 regulatory subunits; and γ correspond to p101 and p84/87 regulatory subunits. The regulatory subunit has the SH2 domain, which can recognize the intracellular kinase domain (TK) of RTKs and trigger the activation of the catalytic subunit p110.

Figure 1. How to activate PI3K
There are 4 ways to activate PI3K. As shown in the figure above, when the insulin family growth factor activates the corresponding RTKs, p85 can be indirectly activated through the adapter protein such as IRS1 (a); in addition, when other family growth factors and RTKs are combined, p85 can also be directly activated. Recognize the TK domain (b); in the form of RAS-GTP, the RAS protein we mentioned in the previous issue can also directly activate the catalytic subunit p110 of PI3K (c). Both Raf and PI3K can be regulated by RAS, which is the main reason why the MAPK pathway and the PI3K pathway overlap; in addition, the γ-type PI3K catalytic subunit can also be activated by some G protein-coupled receptors (d).
02 P13K catalytic function

Figure 2. Reaction catalyzed by PI3K
The activation of PI3K can catalyze the phosphorylation of the 3-hydroxyl of PIP2 to generate PIP3, thereby further promoting downstream pathways; on the contrary, another enzyme in the cell, PTEN, can catalyze the reverse process of this reaction and inhibit the downstream pathways of PI3K.
After PIP3 is generated, it can act as a second messenger, recruiting PDK1 and AKT proteins to the plasma membrane at the same time, so that PDK1 phosphorylates threonine (T308) at position 308 of AKT protein, leading to partial activation of AKT. The activated AKT will further activate downstream regulatory pathways.
03 PI3k-AKT pathway

Figure 3. PI3K-AKT and downstream proteins
AKT is a type of serine/threonine kinase of the AGC family, which mainly contains three domains: PH domain (has affinity for PIP3 and is essential for binding to the cell membrane), catalytic domain, and regulatory domain. As a proto-oncogene, AKT plays a key role in various cellular processes, such as glucose metabolism, cell apoptosis, cell proliferation, transcription and cell migration; PI3K/AKT signaling pathway dysregulation is seen in many human diseases, including cancer and diabetes , Cardiovascular disease and neurological disease.
There are many types of substrates downstream of AKT. In combination with the above figure, we will summarize the main substrates:
1) TSC2
TSC complex (tuberous sclerosis complex) contains three subunits: TSC1, TSC2 and TBC1D7, which can negatively regulate the activity of mTORC1 complex and play a role in tumor suppression. The main principle for this effect is: the GTPase activating protein (GAP) domain contained in TSC2 hydrolyzes the GTP form of RHEB (a small G protein in the RAS family) into the GDP form of RHEB, resulting in RHEB is inactivated and cannot activate the kinase activity of mTORC1, thereby inhibiting cell growth. When AKT induces phosphorylation of Ser 939 and Thr of 1462 of TSC2 protein (tuberin), the activity is inhibited, and the mTOR pathway is activated instead;
2) BAD
BAD (Bcl2-related cell death agonist) can positively regulate mitochondrial apoptosis. After phosphorylation of this protein by AKT, BAD activity is inhibited;
3) MDM2
AKT can activate the ubiquitin ligase-MDM2 of p53 by phosphorylation, thereby inhibiting cell cycle arrest and apoptosis caused by p53.
4) p21 and p27
CDK inhibitors p21 and p27 are two important cell cycle regulators. When normal cell DNA is damaged, they are induced by P53 to inhibit DNA replication, enhance DNA repair, and play a role in suppressing cancer. The phosphorylation of AKT can inhibit the activity of these CDK inhibitors.
5) GSK3β
GSK3β (glycogen synthase kinase 3β) is a very conserved serine/threonine kinase in evolution. It is involved in glycogen metabolism and plays an important role in the Wnt pathway. Under normal circumstances, GSK3β can phosphorylate β-catenin and make it degraded by the proteasome. After AKT phosphorylates GSK3β, GSK3β is inactivated, and β-catenin cannot be phosphorylated again, causing β-catenin to accumulate in the cytoplasm and enter the nucleus. And activate genes related to cell division and growth regulation (such as c-myc and Cyclin D1, etc.).
6) IKK (IκB kinase)
IKK (IκB kinase) is activated by phosphorylation of AKT. IKK further phosphorylation inhibits the transcription factor IκB, which can inhibit the NF-kβ pathway (pay attention to the sentence…). The final result is: AKT positively regulates the NF-kβ pathway, thereby It has an impact on cell survival, proliferation, invasion, angiogenesis and chemotherapy resistance.
7) WEE1
CDK1 inhibits kinase-WEE1 can be inhibited by phosphorylation, which has an effect on mitosis.
8) ASK1
ASK1 (apoptotic signal-regulated kinase 1) is a MAPKKK that can activate the JNK signaling pathway (a type of MAPK pathway, see the second phase for details), and is related to apoptosis induction, pro-apoptosis and endothelial reticulum stress (ERS) After AKT phosphorylates ASK1, its activity is inhibited.
9) FOXO
FOXO is an apoptosis-inducing transcription factor. Phosphorylation of FOXO by AKT causes it to relocate in the cell and lose its transcriptional activity.
In summary, we can see that the effects produced by AKT eventually lead to cell proliferation and avoid apoptosis.
04 PI3K-AKT-mTOR pathway and TORC complex
As we have said before, AKT can indirectly lead to the enhancement of mTOR activation by RHEB by inhibiting TSC2. The full name of mTOR is called the mammalian target of rapamycin (mTOR). mTOR is a type of serine/threonine kinase, the C-terminus is homologous to the catalytic domain of phosphatidylinositol kinase (PI3K), but it does not have the activity of esterase kinase, but has Ser/Thr protein kinase activity. mTOR can form two complexes: TORC1 and TORC2.

Figure 4. PI3K-AKT-mTOR pathway
In addition to being activated by AKT, mTOR can also be activated or inhibited according to the energy and nutrition of the cellular environment. As shown in the figure above, when energy is insufficient (the AMP/ATP ratio increases) and hypoxia, the formation of TSC2 and TSC1 complexes is enhanced and the formation of mTOR complexes is inhibited. When there is an excess of lysosomal amino acids, H+-vATPase can activate Ragulator, a protein complex with guanine nucleotide exchange factors (GEFs), thereby promoting the formation of TORC1.
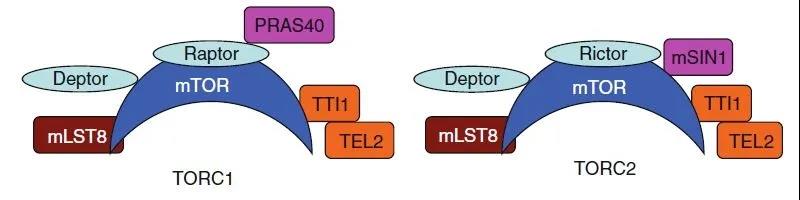
Figure 5. TORC1 and TORC2
As mentioned earlier, mTOR has two complexes: mTORC1 and mTORC2. mTORC1 is composed of mTOR, mTOR regulatory related proteins Raptor, MLST8, and non-core components PRAS40 and DEPTOR. mTORC2 is composed of mTOR, RICTOR, MLST8 and mSIN1. The main difference between the two complexes is: whether the binding protein is Raptor or Rictor and whether the related protein is PRAS40 or mSIN1. Among them, the difference of the former mainly determines that TORC1 is sensitive to rapamycin, while TORC2 is insensitive to rapamycin; the difference of the latter is that: PRAS40 is the substrate of AKT, and mSIN1 is the substrate of MAPK.
From a functional point of view, TORC2 has a relatively simple function. It can phosphorylate AKT to generate positive feedback on mTOR and promote further activation of mTOR (echoing the partial activation of PDK1 in the previous section). TORC1 is different. It can continue to phosphorylate a variety of downstream substrates and regulate biological processes such as cell survival, proliferation, protein synthesis, lipid production, angiogenesis, and autophagy.
05 TORC1 downstream substrates and effects

Figure 6. Substrates and effects downstream of TORC1
TORC1 and protein synthesis
Eukaryotic initiation factor 4E (eIF4E) is a cap-binding protein that can specifically recognize the cap structure at the 5’end of mRNA and play an important role in the initiation of eukaryotic translation. When the repressor protein 4EBP1 is phosphorylated, eIF4E cannot be released to recruit ribosomes to activate the translation process. mTOR relieves its inhibitory effect on EIF4E by phosphorylating 4EBP1, thereby promoting translation and protein synthesis.
TORC1 can also phosphorylate and activate the downstream AGC family ribosomal S6 kinase (S6K1). S6K1 can phosphorylate and activate ribosomal S6 protein (RPS6), PDK1 (AKT upstream agonist protein), MDM2 (p53 inhibitor), transcription regulator EEF2 , Adaptor protein IRS, etc., phosphorylation inhibits the pro-apoptotic protein BAD. The final effect is: avoid cell apoptosis and promote protein synthesis.
TORC1 and lipid synthesis
In addition to promoting protein synthesis, TORC1 can also phosphorylate and activate STAT3 (signal transducer and activator of transcription 3) to promote cell proliferation; activate SREBPs (sterol regulatory element binding proteins) and PPARα and γ (peroxisome proliferator activation) Receptor) promote lipid synthesis.
TORC1 and angiogenesis
TORC1 can promote angiogenesis by activating HIF1α, and HIF1α can also indirectly activate TSC2, thereby negatively regulating TORC.
TORC1 and autophagy
TORC1 inhibits the downstream substrate ULK1 and other autophagy-related proteins through phosphorylation, and inhibits the autophagy process.
In summary, TORC1 activates/inhibits a series of substrates through phosphorylation, directly/indirectly increases the synthesis of proteins and lipids, promotes cell proliferation, and causes the growth of cells and tissues.
06 PI3K-AKT-mTOR and tumor
From the above we know that the PI3K pathway can avoid cell apoptosis and promote its survival and proliferation in a variety of ways. PI3K/AKT signaling pathway is widely activated in tumorigenesis, especially PIK3CA, PIK3R1, PTEN, AKT and other genes have high-frequency mutations (including PIK3CA gene mutations in approximately 36% of breast cancers), which is related to tumorigenesis, development, and Resistance is closely related.
At present, the main mutations involved in this pathway are:
1) PIK3CA mutation
PIK3CA is a gene encoding the P110 subunit of PI3K protein in tumors and has a high mutation rate. In many cancers such as prostate cancer and breast cancer, more than 10% of cases have PIK3CA mutations. Among them, activating mutations account for more than 80%. The activation point mutation allows the PI3K protein to remain activated without relying on upstream signals. PIK3CA mutation is considered to be one of the potential causes of resistance to EGFR-TKI therapy.
2) PTEN mutation
As we mentioned earlier, the tumor suppressor gene PTEN can produce the opposite effect of PI3K, so its mutations also lead to cancer development. In endometrial cancer, brain cancer, skin cancer, and prostate cancer, more than 10% of cases have PTEN gene mutations.
3) AKT mutation
The AKT gene family includes: AKT1, AKT2, and AKT3. The most common forms of AKT mutations in tumors are activation point mutations and abnormal increase in gene copy number. In breast cancer, colorectal cancer, ovarian cancer and bladder cancer, there are quite a high proportion of cases.

Figure 7. Drugs designed for PI3K-AKT-mTOR pathway
As shown in the figure above, the current drug development for PI3K, AKT and mTORC targets has some progress.
(source:internet, reference only)
Disclaimer of medicaltrend.org



