mRNA: Optimization and Application in diseases and cancer
mRNA: Optimization and Application in diseases and cancer
- Should China be held legally responsible for the US’s $18 trillion COVID losses?
- CT Radiation Exposure Linked to Blood Cancer in Children and Adolescents
- Can people with high blood pressure eat peanuts?
- What is the difference between dopamine and dobutamine?
- What is the difference between Atorvastatin and Rosuvastatin?
- How long can the patient live after heart stent surgery?
mRNA: Optimization and Application in diseases and cancer.
Messenger RNA (mRNA)-based drugs, especially mRNA vaccines, have been widely proven to be promising therapeutic strategies in immunotherapy.
Some special advantages of mRNA vaccines, including: high efficacy, low cost, low side effects, etc., make it popular in preclinical and clinical trials for various infectious diseases and cancer.
Some recent technological advances have also alleviated the problems faced in the development of mRNA vaccines and greatly improved the immunogenicity of mRNA vaccines.
This article reviews the optimization methods of mRNA vaccines, and the potential biological mechanisms of this method, and introduces the applications of mRNA vaccines in infectious diseases and cancer.
1. Introduction to mRNA vaccines
The messenger ribonucleic acid mRNA is an intermediate genetic material in the central law. It was first discovered in 1961 [1], and in 1989 the concept of mRNA-based drugs was proposed [2].
In 1990, in vitro transcribed mRNA was fully expressed in mouse bone marrow muscle cells by direct injection, proving the feasibility of mRNA vaccine development (Figure 1).
As a result, related technologies such as mRNA structure research have developed rapidly, alleviating several limitations caused by mRNA instability, high innate immunogenicity and low delivery efficiency in vivo.
The current mRNA vaccines have been extensively studied in different types of diseases.
 Figure 1: Some key findings and progress schedule based on mRNA technology
Figure 1: Some key findings and progress schedule based on mRNA technology
2. Special advantages of mRNA vaccines
mRNA vaccines show many special advantages that traditional vaccines do not have.
First, mRNA can theoretically satisfy all the genetic information needed to encode and express various proteins.
Second, by modifying the mRNA sequence, the efficiency of vaccine development can also be optimized. Such optimization is relatively simpler than other vaccines, and may be retained or standardized by different antigens.
Third, mRNA has self-assisting properties. It activates a strong and long-lasting adaptive immune response through tumor necrosis factor-α (TNF-α), interferon-α (IFN-α) and other cytokines secreted by immune cells (Figure 2 ), while peptide and protein-based vaccines require additional adjuvants to achieve similar goals.
The in vivo expression of mRNA can also avoid protein and virus-derived pollution. Fourth, compared with DNA-based vaccines, mRNA vaccines can express target proteins more effectively, because they are expressed in the cytoplasm without entering the nucleus, are less likely to integrate into the host DNA genome, and induce less immunity Rejection.
In addition, mRNA only has a short-term activity and is easily completely decomposed through physiological metabolic pathways. Therefore, it will not become a burden for the host’s homeostasis.
 Figure 2: In vitro transcription of mRNA and innate immune activation
Figure 2: In vitro transcription of mRNA and innate immune activation
3. In vitro synthesis and optimization of mRNA
So far, the in vitro transcription technology of mRNA has been mature, and the most commonly used method is to use T3, T7 or SP6 RNA polymerase and linear DNA (linearized plasmid DNA or synthetic DNA prepared by PCR) for mRNA synthesis.
Complete functional mRNA has some basic structural elements, including 5’cap, 5’UTR, open reading frame (ORF), 3’UTR and poly(A) tail, which keeps the mRNA structure intact and is conducive to the stability of mRNA Sexuality and expressive ability.
(1) 5’ hat optimization
Capping with cap analogs is the most common capping method for in vitro transcription of mRNA.
However, studies have found that conventional cap analogs can bind mRNA sequences in reverse. In order to avoid reverse incorporation of 5’caps, reverse cap analogs have been developed (ARCA), ARCA is modified at the C2 or C3 position, which has higher translation efficiency. Another cap analog was developed in 2018, a co-transcription capping method called “CleanCap”.
It uses the trimer that initiates the end-capping to produce the naturally occurring 5’cap structure, which increases the end-capping efficiency to nearly 90-99%.
(2) UTR optimization
UTR is the non-coding part of the upstream and downstream structural domai
ns located in the mRNA coding region. It is related to the process of mRNA replication and translation. In order to improve the stability and translation efficiency of mRNA, the optimization of UTRs must be ensured.
5’UTR directly affects the translation of the downstream ORF, avoiding that the gene sequence in the 5’UTR is the same as the upstream of the ORF, which can effectively prevent the wrong start and replacement of the reading frame during the mRNA translation process, and some specific sequences can also be added to the 5′ UTR to enhance the stability of mRNA and the accuracy of translation. For example, Kozak et al. inserted the sequence GCC-(A/G)-CCAUGG in this region to start the translation process more accurately.
Stability regulatory elements are often added to 3’UTR. For example, the widely used 3’UTR sequences derived from α-globin and β-globin contain translation and stability regulatory elements.
Avoiding unstable sequences when synthesizing 3’UTR can increase mRNA stability, and introducing stable elements into 3’UTR can also significantly improve mRNA stability and extend its half-life.
(3) ORF optimization
Choosing appropriate codons in the ORF region can optimize the overall translation efficiency of mRNA.
The optimized ORF sequence usually contains synonymous frequent codons with higher tRNA abundance and/or codons to replace rare codons in the ORF.
Use the same codons of the host to translate, highly express genes and/or ensure the integrity of tRNA during expression.
(4) Poly(A) tail optimization
Poly(A) sequence can slow down the degradation process of RNA exonuclease, increase stability, extend the half-life in vivo, and improve the translation efficiency of mRNA.
In addition, Poly(A) binding protein (PABP) can be connected to the 5’cap through translation initiation factors (such as elF4G and elF4E), thereby affecting the closed-loop structure of mRNA and coordinating the stability and translation efficiency of mRNA.
Poly(A) sequences of different lengths can affect the translation efficiency of mRNA to varying degrees.
Since the length of the Poly(A) sequence required for high translation efficiency mRNA in various types of cells is different,
adjustments should be made to optimize the translation efficiency of mRNA.
(5) Optimization of delivery system
There are some difficulties when mRNA enters the host cytoplasm. First, the mRNA molecules are not small enough to diffuse freely through the cell membrane. Second, both the mRNA and the cell membrane are negatively charged, which increases the difficulty of mRNA delivery. Third, the mRNA is easily affected by the skin and The extracellular ribonuclease present in the blood is degraded. Therefore, delivering mRNA to a sufficient number of cells with a sufficiently high translation level is one of the most difficult application problems for mRNA vaccines, because it requires a highly specific and effective mRNA delivery system (Figure 3).


Figure 3: mRNA delivery system
4. mRNA vaccine clinical research
Immunotherapy, especially vaccines against infectious diseases and cancer, is the core area of the mRNA drug platform.
(1) mRNA vaccine for infectious diseases
The mRNA vaccines for infectious diseases have made promising achievements, and some products have entered the stage of human clinical trials (Figure 4).
The overall development steps of these vaccines are:
(1) constructing a core antigen-encoding mRNA sequence optimized or combined based on selected antigens from target pathogens;
(2) trying and selecting mRNA construction types, adjuvants, carrier materials and administration routes;
( 3) Detect the in vivo expression of the encoded antigen and the level of immune induction mechanism;
(4) Provide research and proof of immune induction mechanism.
 Figure 4: Clinical trial of infectious disease mRNA vaccine
Figure 4: Clinical trial of infectious disease mRNA vaccine
(2) mRNA tumor vaccine
Tumor-associated antigens and tumor-specific new epitopes are the core targets of mRNA tumor vaccines.
Tumor vaccines are now aimed at stimulating cellular immunity to treat the diversity and uncertainty of tumor occurrence. Some related clinical trials are underway (Figure 5).
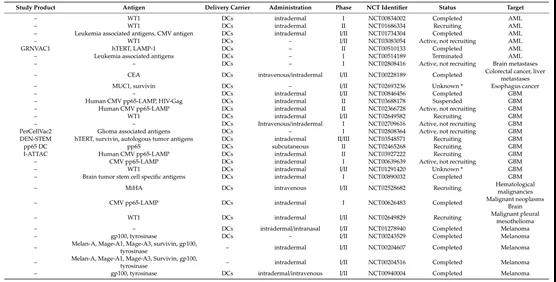

Figure 5: mRNA vaccines used in cancer clinical trials
Summary
The mRNA vaccine is a powerful and multifunctional platform, and its clinical success will significantly enhance our ability to respond to and control emerging infectious diseases.
However, we still need to further study the transport of mRNA outside the cell and the expression and translation inside the cell.
In the next few years, several key clinical trials of mRNA vaccines are about to be completed (especially for COVID-19), and broader clinical experience will give us a more comprehensive understanding of mRNA vaccine platforms, thereby improving mRNA and various The production capacity of carrier materials, screening of potential carrier molecules and adjuvants, provide a basis for the industrialization of mRNA vaccines.
Although the field of mRNA vaccines is still far from mature, its potential as a preferred vaccine model has been fully demonstrated.
mRNA: Optimization and Application in diseases and cancer
(source:internet, reference only)
Disclaimer of medicaltrend.org
Important Note: The information provided is for informational purposes only and should not be considered as medical advice.



