Science: Nanopores can directly sequence proteins with 100% accuracy
- Normal Liver Cells Found to Promote Cancer Metastasis to the Liver
- Nearly 80% Complete Remission: Breakthrough in ADC Anti-Tumor Treatment
- Vaccination Against Common Diseases May Prevent Dementia!
- New Alzheimer’s Disease (AD) Diagnosis and Staging Criteria
- Breakthrough in Alzheimer’s Disease: New Nasal Spray Halts Cognitive Decline by Targeting Toxic Protein
- Can the Tap Water at the Paris Olympics be Drunk Directly?
- Should China be held legally responsible for the US’s $18 trillion COVID losses?
- CT Radiation Exposure Linked to Blood Cancer in Children and Adolescents
- FDA has mandated a top-level black box warning for all marketed CAR-T therapies
- Can people with high blood pressure eat peanuts?
- What is the difference between dopamine and dobutamine?
- How long can the patient live after heart stent surgery?
Science: Nanopores can directly sequence proteins with 100% accuracy and can also identify amino acid modifications
Protein is the main component of living organisms, and it is also the main undertaker of life activities. Proteins with biological functions often have specific spatial structures, and protein structures are defined on multiple levels.
Among them, the primary structure, that is, the type and arrangement of amino acids, is the most important, and it can determine the high-level structure of the protein. However, it has always been very difficult to directly read the primary structure of a protein .
In most cases, scientists will “decipher” the amino acid sequence of a protein based on the gene sequence and amino acid codon table. However, due to the existence of post-transcriptional modification and post-translational modification, the deciphering result is not completely correct, and it is even quite different from the real amino acid sequence.
On November 4, 2021, researchers from Delft University of Technology in the Netherlands published a titled Multiple rereads of single proteins at single-amino acid resolution using nanopores in Science .
Research papers on a single protein with multiple rereads). The study used nanopore sequencing technology to successfully scan and read the amino acid sequence of a single protein : linearized DNA-peptide complexes slowly pass through a tiny nanopore, and researchers can read related proteins according to the change and intensity of the current Information content, directly sequence the amino acid sequence of the protein .

Protein is the main bearer of life activities. In fact, all biological proteins are long peptide chains composed of about 20 different amino acids, just like there are different kinds of beads on a necklace. Unfortunately, current protein sequencing methods are expensive and cannot detect many rare proteins. The nanopore sequencing technology developed in recent years has been able to directly scan and sequence single DNA molecules.
Today, this article is published in ScienceThe above research shows that we can directly read the amino acid sequence of proteins in a manner similar to DNA nanopore sequencing.
Professor Cees Dekker , the corresponding author of this study, said: In the past 30 years, DNA sequencing based on nanopores has evolved from an idea to a practical working device, and has successfully developed a commercial portable nanopore sequencer to serve The billion-dollar genome sequencing market. In our paper, we extend the concept of nanopores to the reading of individual proteins.
This may have a major impact on basic protein research and medical diagnosis.

Oxford nanopores developed nanopore sequencing instrument directly read the amino acid sequence features of the nanopore how to use a single amino acid peptide chain read, the first author of the paper Henry Brinkerhoff doctor made a vivid metaphor: “Imagine Now, the amino acid chain in a peptide chain is like a necklace with beads of different sizes.
Then, you turn on the faucet and slowly send the necklace into the sewer, which is the nanopore. If at a certain point in time it is a Large beads, it will block the sewer, and the water inside will become a trickle. On the contrary, if it is a small bead, then the remaining gap in the sewer will be larger and the water flow will be larger.”
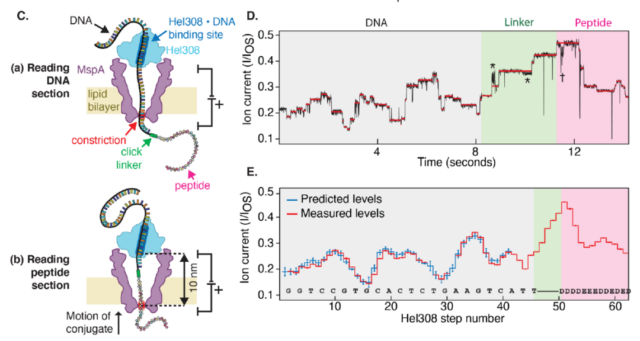
Read with Nanopore Peptide The device directly reads the amino acid sequence. Therefore, through this technology, researchers can very accurately measure the current size of the nanopore, and use this to predict the corresponding amino acid species.
More importantly, this process does not affect the integrity of the peptide chain, so we can read a single peptide chain again and again, and then fit all the data, so as to obtain the peptide chain with basically 100% accuracy. The sequence composition.
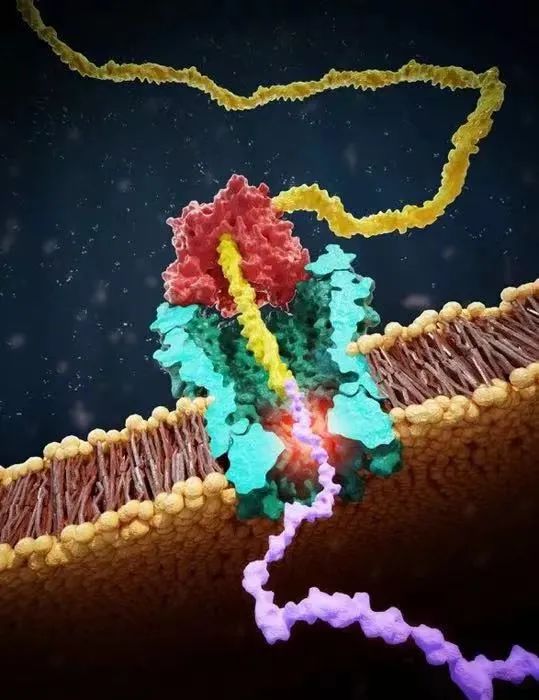
The helicase (red) drags the DNA molecule (yellow) connected with the polypeptide (purple) to slowly pass through the nanopore (green), thereby characterizing the amino acid sequence of the polypeptide by reading the electrical signal (highlighted in orange).
Barcode-like recognition accuracy To further verify the accuracy of this technology, the researchers changed a certain amino acid of the peptide chain, and then were able to detect significantly different electrical signals, indicating that the technology is extremely sensitive.
In fact, this new technology is very powerful in identifying individual proteins and mapping the subtle changes between them, as an analogy—just like a cashier in a supermarket scans a barcode to identify each product. This may also provide a new way for de novo protein sequencing in the future.
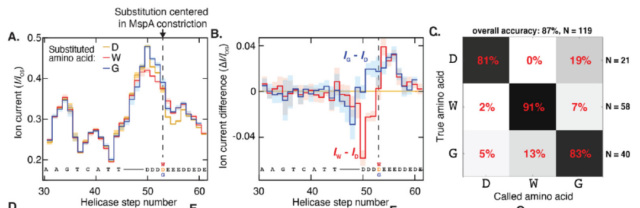
The nanopore peptide reader can distinguish single peptides substituted by single amino acids , Dr. Henry Brinkerhoff said: This method may lay the foundation for future protein sequencing, but for now, de novo protein sequencing is still a huge challenge.
We still need a large number of descriptions of electrical signals from different sequences in order to create a ” code table ” corresponding to electrical signals and protein sequences . But even so, the research has been able to successfully distinguish individual amino acid changes in the protein sequence.
This is undoubtedly a major advancement and will have many direct applications. Seeing the biological “dark matter” Dark matter is an invisible matter that may exist in the universe theoretically.
It may be the main component of the universe’s matter, but it does not belong to any known form of visible celestial bodies. Of the substance. In cells, there are many unknowable ” dark matter “-millions of protein mutations caused by post-translational modifications.
These protein mutations cannot be predicted by gene sequences, but the emergence of nanoporin sequencing technology has reversed this situation. Using the current nanopore peptide reader, researchers can directly observe these “dark matter in biology.”
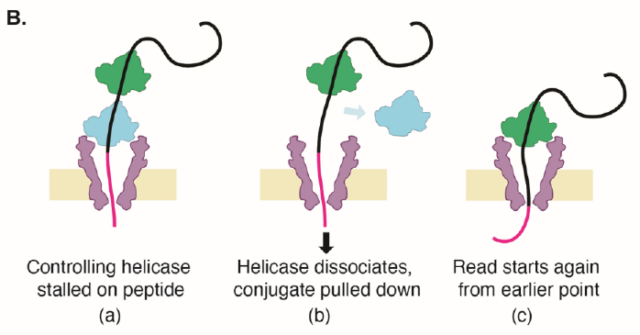
Reading a single protein repeatedly to improve the accuracy is easy to understand. The corresponding author, Professor Cees Dekker , made an analogy: the beads on the necklace are not only different in size, but also in different colors. For example, some red beads represent a phosphate group attached. The other blue beads represent a sugar group attached.
These changes are critical to protein function and are also a sign of diseases such as cancer. Our new method will be able to detect these changes, thereby providing a new theoretical basis for the detection and treatment of diseases such as cancer .
Conclusion In conclusion, this proteomics tool that can directly read protein sequences is of great significance to the research and application of cell biology.
The study demonstrated a single-molecule peptide reader based on nanopores, which uses DNA helicase Hel308 to pass DNA-peptide conjugates through the biological nanopore MspA, and reads the amino acid sequence of linearized proteins according to changes in current.
More importantly, this method can distinguish individual amino acid changes, with high fidelity and high throughput potential.
This single-molecule peptide reader marks a new breakthrough in protein identification and opens the way for single-molecule protein sequencing and classification in single cells.
Link to the paper: https://www.science.org/doi/10.1126/science.abl4381
Science: Nanopores can directly sequence proteins with 100% accuracy and can also identify amino acid modifications
(source:internet, reference only)
Disclaimer of medicaltrend.org
Important Note: The information provided is for informational purposes only and should not be considered as medical advice.



